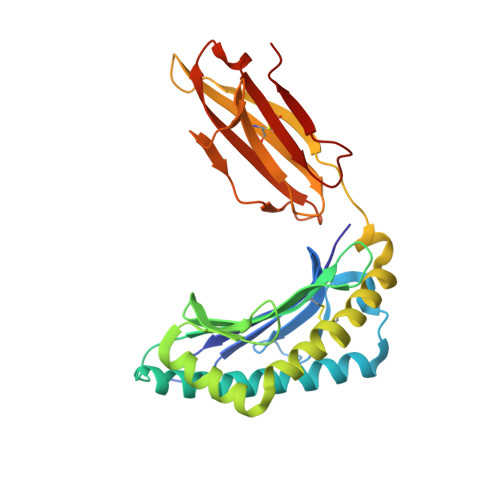
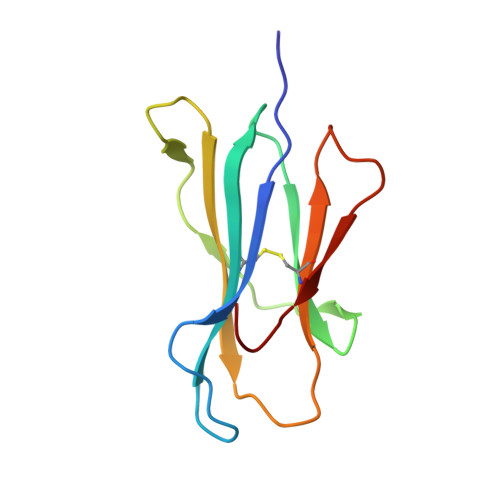
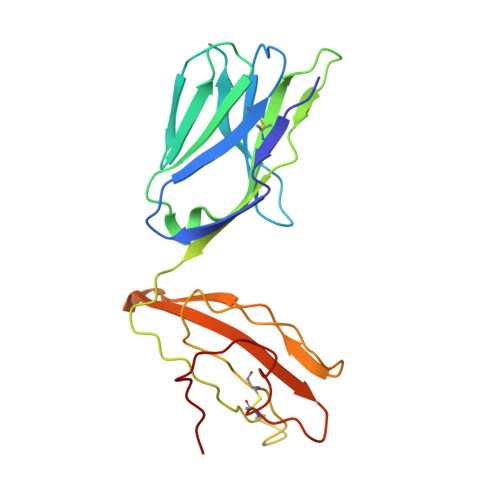
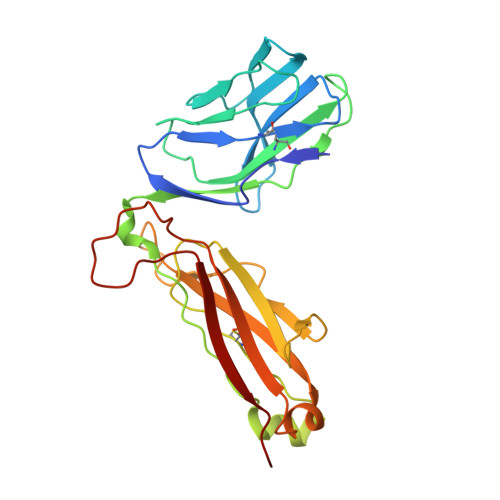
The Structural Dynamics and Energetics of an Immunodominant T-Cell Receptor are Programmed by its Vbeta Domain
Ishizuka, J., Stewart-Jones, G., Van Der Merwe, A., Bell, J., Mcmichael, A., Jones, Y.(2008) Immunity 28: 171
- PubMed: 18275829 Search on PubMed
- DOI: https://doi.org/10.1016/j.immuni.2007.12.018
- Primary Citation Related Structures:
2VLJ, 2VLK, 2VLL, 2VLM, 2VLR - PubMed Abstract:
Immunodominant and public T cell receptor (TCR) usage is relatively common in many viral diseases yet surprising in the context of the large naive TCR repertoire. We examined the highly conserved Vbeta17:Valpha10.2 JM22 T cell response to the influenza matrix peptide (58-66)-HLA-A*0201 (HLA-A2-flu) through extensive kinetic, thermodynamic, and structural analyses. We found several conformational adjustments that accompany JM22-HLA-A2-flu binding and identified a binding "hotspot" within the Vbeta domain of the TCR. Within this hotspot, key germline-encoded CDR1 and CDR2 loop residues and a crucial but commonly coded residue in the hypervariable region of CDR3 provide the basis for the substantial bias in the selection of the germline-encoded Vbeta17 domain. The chances of having a substantial number of T cells in the naive repertoire that have HLA-A2-flu-specific Vbeta17 receptors may consequently be relatively high, thus explaining the immunodominant usage of this clonotype.
- MRC Human Immunology Unit, Weatherall Institute of Molecular Medicine, University of Oxford, John Radcliffe Hospital, Oxford, UK.
Organizational Affiliation: