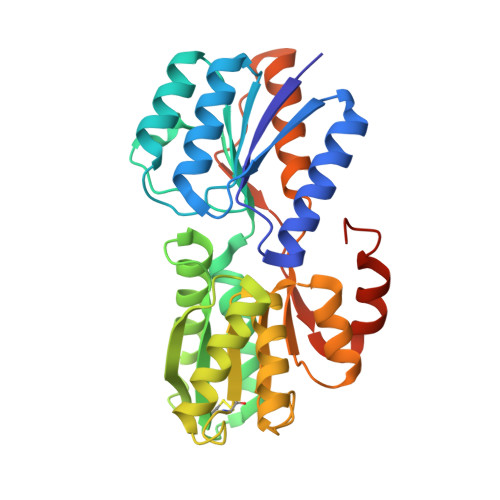
Furanose-Specific Sugar Transport: Characterization of a Bacterial Galactofuranose-Binding Protein.
Horler, R.S.P., Muller, A., Williamson, D.C., Potts, J.R., Wilson, K.S., Thomas, G.H.(2009) J Biological Chem 284: 31156
- PubMed: 19744923 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.054296
- Primary Citation Related Structures:
2VK2 - PubMed Abstract:
The widespread utilization of sugars by microbes is reflected in the diversity and multiplicity of cellular transporters used to acquire these compounds from the environment. The model bacterium Escherichia coli has numerous transporters that allow it to take up hexoses and pentoses, which recognize the more abundant pyranose forms of these sugars. Here we report the biochemical and structural characterization of a transporter protein YtfQ from E. coli that forms part of an uncharacterized ABC transporter system. Remarkably the crystal structure of this protein, solved to 1.2 A using x-ray crystallography, revealed that YtfQ binds a single molecule of galactofuranose in its ligand binding pocket. Selective binding of galactofuranose over galactopyranose was also observed using NMR methods that determined the form of the sugar released from the protein. The pattern of expression of the ytfQRTyjfF operon encoding this transporter mirrors that of the high affinity galactopyranose transporter of E. coli, suggesting that this bacterium has evolved complementary transporters that enable it to use all the available galactose present during carbon limiting conditions.
- Department of Biology, University of York, York YO10 5YW, United Kingdom.
Organizational Affiliation: