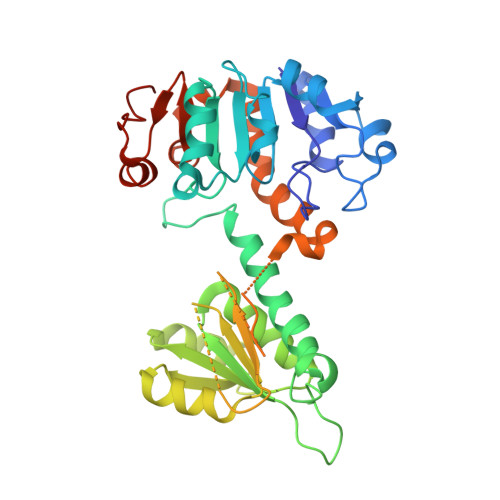
Three-Dimensional Structures of Apo- and Holo-L-Alanine Dehydrogenase from Mycobacterium Tuberculosis Reveal Conformational Changes Upon Coenzyme Binding.
Agren, D., Stehr, M., Berthold, C.L., Kapoor, S., Oehlmann, W., Singh, M., Schneider, G.(2008) J Mol Biology 377: 1161
- PubMed: 18304579 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.091
- Primary Citation Related Structures:
2VHV, 2VHW, 2VHX, 2VHY, 2VHZ - PubMed Abstract:
L-alanine dehydrogenase from Mycobacterium tuberculosis catalyzes the NADH-dependent reversible conversion of pyruvate and ammonia to L-alanine. Expression of the gene coding for this enzyme is up-regulated in the persistent phase of the organism, and alanine dehydrogenase is therefore a potential target for pathogen control by antibacterial compounds. We have determined the crystal structures of the apo- and holo-forms of the enzyme to 2.3 and 2.0 A resolution, respectively. The enzyme forms a hexamer of identical subunits, with the NAD-binding domains building up the core of the molecule and the substrate-binding domains located at the apical positions of the hexamer. Coenzyme binding stabilizes a closed conformation where the substrate-binding domains are rotated by about 16 degrees toward the dinucleotide-binding domains, compared to the open structure of the apo-enzyme. In the structure of the abortive ternary complex with NAD+ and pyruvate, the substrates are suitably positioned for hydride transfer between the nicotinamide ring and the C2 carbon atom of the substrate. The approach of the nucleophiles water and ammonia to pyruvate or the reaction intermediate iminopyruvate, respectively, is, however, only possible through conformational changes that make the substrate binding site more accessible. The crystal structures identified the conserved active-site residues His96 and Asp270 as potential acid/base catalysts in the reaction. Amino acid replacements of these residues by site-directed mutagenesis led to inactive mutants, further emphasizing their essential roles in the enzymatic reaction mechanism.
- Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 77 Stockholm, Sweden.
Organizational Affiliation: