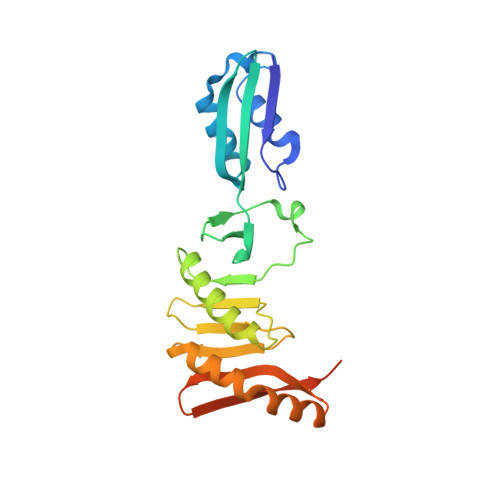
Structural and Mutational Analysis of Cell Division Protein Ftsq
van den Ent, F., Vinkenvleugel, T., Ind, A., West, P., Veprintsev, D., Naninga, N., den Blaauwen, T., Lowe, J.(2008) Mol Microbiol 68: 110
- PubMed: 18312270
- DOI: https://doi.org/10.1111/j.1365-2958.2008.06141.x
- Primary Citation Related Structures:
2VH1, 2VH2 - PubMed Abstract:
Bacterial cytokinesis requires the divisome, a complex of proteins that co-ordinates the invagination of the cytoplasmic membrane, inward growth of the peptidoglycan layer and the outer membrane. Assembly of the cell division proteins is tightly regulated and the order of appearance at the future division site is well organized. FtsQ is a highly conserved component of the divisome among bacteria that have a cell wall, where it plays a central role in the assembly of early and late cell division proteins. Here, we describe the crystal structure of the major, periplasmic domain of FtsQ from Escherichia coli and Yersinia enterocolitica. The crystal structure reveals two domains; the alpha-domain has a striking similarity to polypeptide transport-associated (POTRA) domains and the C-terminal beta-domain forms an extended beta-sheet overlaid by two, slightly curved alpha-helices. Mutagenesis experiments demonstrate that two functions of FtsQ, localization and recruitment, occur in two separate domains. Proteins that localize FtsQ need the second beta-strand of the POTRA domain and those that are recruited by FtsQ, like FtsL/FtsB, require the surface formed by the tip of the last alpha-helix and the two C-terminal beta-strands. Both domains act together to accomplish the role of FtsQ in linking upstream and downstream cell division proteins within the divisome.
- MRC-LMB, Hills Road, Cambridge CB2 2QH, UK. fent@mrc-lmb.cam.ac.uk
Organizational Affiliation: