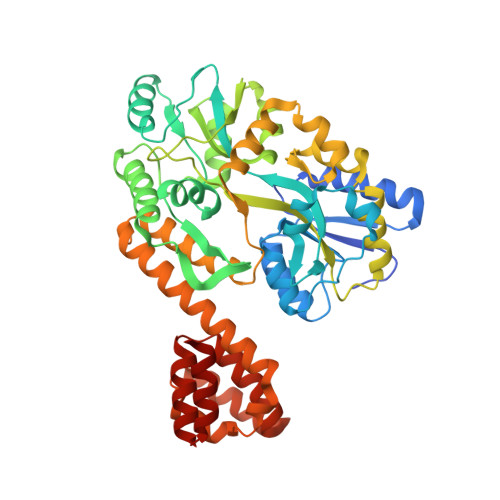
Crystal Structure of Human Ips-1 Caspase Activation Recruitment Domain
Potter, J.A., Randall, R.E., Taylor, G.L.(2008) BMC Struct Biol 8: 11
- PubMed: 18307765 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-8-11
- Primary Citation Related Structures:
2VGQ - PubMed Abstract:
IPS-1/MAVS/VISA/Cardif is an adaptor protein that plays a crucial role in the induction of interferons in response to viral infection. In the initial stage of the intracellular antiviral response two RNA helicases, retinoic acid inducible gene-I (RIG-I) and melanoma differentiation-association gene 5 (MDA5), are independently able to bind viral RNA in the cytoplasm. The 62 kDa protein IPS-1/MAVS/VISA/Cardif contains an N-terminal caspase activation and recruitment (CARD) domain that associates with the CARD regions of RIG-I and MDA5, ultimately leading to the induction of type I interferons. As a first step towards understanding the molecular basis of this important adaptor protein we have undertaken structural studies of the IPS-1 MAVS/VISA/Cardif CARD region. The crystal structure of human IPS-1/MAVS/VISA/Cardif CARD has been determined to 2.1A resolution. The protein was expressed and crystallized as a maltose-binding protein (MBP) fusion protein. The MBP and IPS-1 components each form a distinct domain within the structure. IPS-1/MAVS/VISA/Cardif CARD adopts a characteristic six-helix bundle with a Greek-key topology and, in common with a number of other known CARD structures, contains two major polar surfaces on opposite sides of the molecule. One face has a surface-exposed, disordered tryptophan residue that may explain the poor solubility of untagged expression constructs. The IPS-1/MAVS/VISA/Cardif CARD domain adopts the classic CARD fold with an asymmetric surface charge distribution that is typical of CARD domains involved in homotypic protein-protein interactions. The location of the two polar areas on IPS-1/MAVS/VISA/Cardif CARD suggest possible types of associations that this domain makes with the two CARD domains of MDA5 or RIG-I. The N-terminal CARD domains of RIG-I and MDA5 share greatest sequence similarity with IPS-1/MAVS/VISA/Cardif CARD and this has allowed modelling of their structures. These models show a very different charge profile for the equivalent surfaces compared to IPS-1/MAVS/VISA/Cardif CARD.
- Centre for Biomolecular Sciences, University of St Andrews, St Andrews, Fife, KY16 9ST, UK. jap7@st-andrews.ac.uk
Organizational Affiliation: