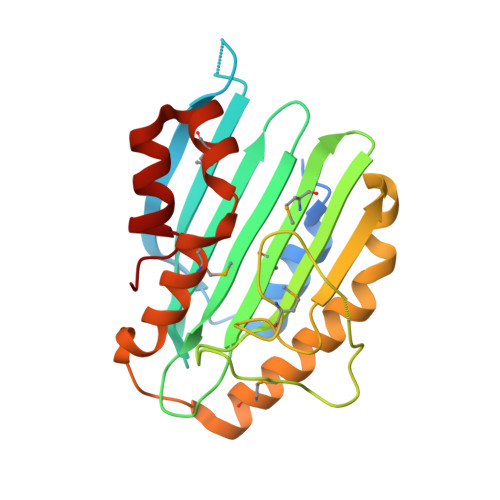
Phycoerythrobilin Synthase (Pebs) of a Marine Virus: Crystal Structures of the Biliverdin Complex and the Substrate-Free Form.
Dammeyer, T., Hofmann, E., Frankenberg-Dinkel, N.(2008) J Biological Chem 283: 27547
- PubMed: 18662988
- DOI: https://doi.org/10.1074/jbc.M803765200
- Primary Citation Related Structures:
2VCK, 2VCL, 2VGR - PubMed Abstract:
The reddish purple open chain tetrapyrrole pigment phycoerythrobilin (PEB; A(lambdamax) approximately 550 nm) is an essential chromophore of the light-harvesting phycobiliproteins of most cyanobacteria, red algae, and cryptomonads. The enzyme phycoerythrobilin synthase (PebS), recently discovered in a marine virus infecting oceanic cyanobacteria of the genus Prochlorococcus (cyanophage PSSM-2), is a new member of the ferredoxin-dependent bilin reductase (FDBR) family. In a formal four-electron reduction, the substrate biliverdin IXalpha is reduced to yield 3Z-PEB, a reaction that commonly requires the action of two individual FDBRs. The first reaction catalyzed by PebS is the reduction of the 15,16-methine bridge of the biliverdin IXalpha tetrapyrrole system. This reaction is exclusive to PEB biosynthetic enzymes. The second reduction site is the A-ring 2,3,3(1),3(2)-diene system, the most common target of FDBRs. Here, we present the first crystal structures of a PEB biosynthetic enzyme. Structures of the substrate complex were solved at 1.8- and 2.1-A resolution and of the substrate-free form at 1.55-A resolution. The overall folding revealed an alpha/beta/alpha-sandwich with similarity to the structure of phycocyanobilin:ferredoxin oxidoreductase (PcyA). The substrate-binding site is located between the central beta-sheet and C-terminal alpha-helices. Eight refined molecules with bound substrate, from two different crystal forms, revealed a high flexibility of the substrate-binding pocket. The substrate was found to be either in a planar porphyrin-like conformation or in a helical conformation and is coordinated by a conserved aspartate/asparagine pair from the beta-sheet side. From the alpha-helix side, a conserved highly flexible aspartate/proline pair is involved in substrate binding and presumably catalysis.
- Physiology of Microorganisms, Ruhr-Universität Bochum, D-44780 Bochum, Germany.
Organizational Affiliation: