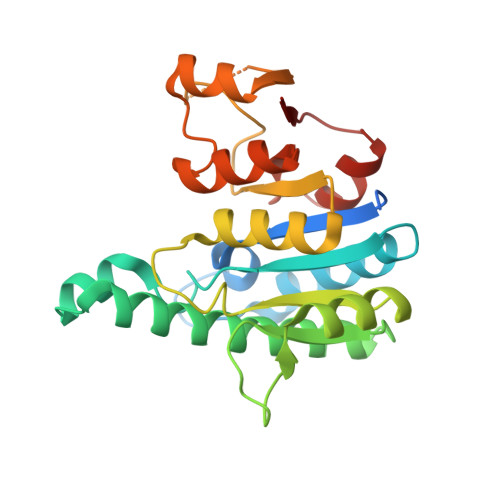
Structural and Functional Investigations of Ureaplasma Parvum Ump Kinase - a Potential Antibacterial Drug Target
Egeblad-Welin, L., Welin, M., Wang, L., Eriksson, S.(2007) FEBS J 274: 6403
- PubMed: 18021254 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-4658.2007.06157.x
- Primary Citation Related Structures:
2VA1 - PubMed Abstract:
The crystal structure of uridine monophosphate kinase (UMP kinase, UMPK) from the opportunistic pathogen Ureaplasma parvum was determined and showed similar three-dimensional fold as other bacterial and archaeal UMPKs that all belong to the amino acid kinase family. Recombinant UpUMPK exhibited Michaelis-Menten kinetics with UMP, with K(m) and V(max) values of 214 +/- 4 microm and 262 +/- 24 micromol.min(-1).mg(-1), respectively, but with ATP as variable substrate the kinetic analysis showed positive cooperativity, with an n value of 1.5 +/- 0.1. The end-product UTP was a competitive inhibitor against UMP and a noncompetitive inhibitor towards ATP. Unlike UMPKs from other bacteria, which are activated by GTP, GTP had no detectable effect on UpUMPK activity. An attempt to create a GTP-activated enzyme was made using site-directed mutagenesis. The mutant enzyme F133N (F133 corresponds to the residue in Escherichia coli that is involved in GTP activation), with F133A as a control, were expressed, purified and characterized. Both enzymes exhibited negative cooperativity with UMP, and GTP had no effect on enzyme activity, demonstrating that F133 is involved in subunit interactions but apparently not in GTP activation. The physiological role of UpUMPK in bacterial nucleic acid synthesis and its potential as target for development of antimicrobial agents are discussed.
- Department of Anatomy, Physiology and Biochemistry, Swedish University of Agricultural Sciences, Uppsala Biomedical Centre, Sweden.
Organizational Affiliation: