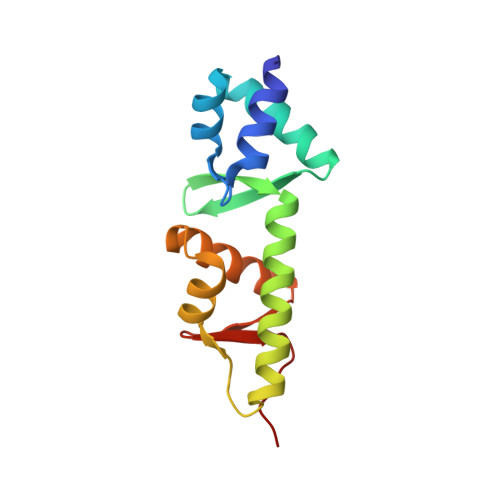
Conformational Switches in Winged-Helix Domains 1 and 2 of Bacterial Translation Elongation Factor Selb.
Ganichkin, O., Wahl, M.C.(2007) Acta Crystallogr D Biol Crystallogr 63: 1075
- PubMed: 17881825 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444907042229
- Primary Citation Related Structures:
2V9V - PubMed Abstract:
The crystal structure of the first two winged-helix motifs of translation elongation factor SelB from Moorella thermoacetica has been determined at 1.1 A resolution. Compared with the previous structure of the two domains in conjunction with winged-helix modules 3 and 4, the first winged-helix domain underwent a substantial conformational change during which the alpha-helical and beta-sheet portions of the element opened up like a shell. This conformational rearrangement was elicited by a change in the orientation of Trp396, leading to the disclosure of a bona fide ligand-binding site in the direct vicinity of Trp396. Additionally, the C-terminal tail of the second domain followed a different path compared with the previous structure. It is conceivable that these conformational switches constitute part of the molecular mechanism that underlies the communication between the N-terminal part of SelB, which binds Sec-tRNA(Sec) and GTP, and the C-terminal part of the protein, which binds selenocysteine-insertion sequences.
- Max-Planck-Institut für Biophysikalische Chemie, Makromolekulare Röntgenkristallographie, Am Fassberg 11, D-37077 Göttingen, Germany.
Organizational Affiliation: