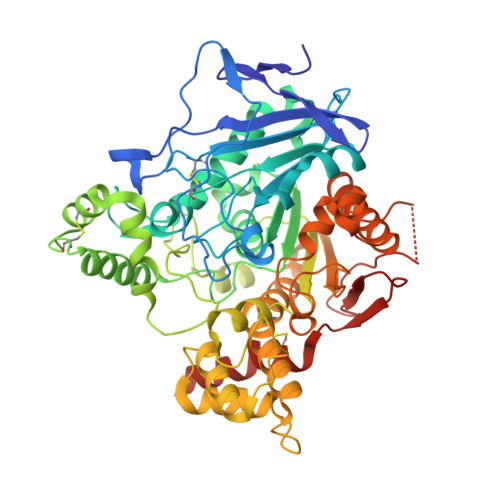
Use of a 'Caged' Analog to Study Traffic of Choline within Acetylcholinesterase by Kinetic Crystallography
Colletier, J.-P., Royant, A., Specht, A., Sanson, B., Nachon, F., Masson, P., Zaccai, G., Sussman, J.L., Goeldner, M., Silman, I., Bourgeois, D., Weik, M.(2007) Acta Crystallogr D Biol Crystallogr 63: 1115
- PubMed: 18007027
- DOI: https://doi.org/10.1107/S0907444907044472
- Primary Citation of Related Structures:
2V96, 2V97, 2V98, 2VA9 - PubMed Abstract:
Acetylcholinesterase plays a crucial role in nerve-impulse transmission at cholinergic synapses. The apparent paradox that it displays high turnover despite its active site being buried raises cogent questions as to how the traffic of substrates and products to and from the active site can occur so rapidly in such circumstances. Here, a kinetic crystallography strategy aimed at structurally addressing the issue of product traffic in acetylcholinesterase is presented, in which UV-laser-induced cleavage of a photolabile precursor of the enzymatic product analogue arsenocholine, 'caged' arsenocholine, is performed in a temperature-controlled X-ray crystallography regime. The 'caged' arsenocholine was shown to bind at both the active and peripheral sites of acetylcholinesterase. UV irradiation of a complex with acetylcholinesterase during a brief temperature excursion from 100 K to room temperature is most likely to have resulted in a decrease in occupancy by the caged compound. Microspectrophotometric experiments showed that the caged compound had indeed been photocleaved. It is proposed that a fraction of the arsenocholine molecules released within the crystal had been expelled from both the active and the peripheral sites. Partial q-weighted difference refinement revealed a relative movement of the two domains in acetylcholinesterase after photolysis and the room-temperature excursion, resulting in an increase in the active-site gorge volume of 30% and 35% in monomers A and B of the asymmetric unit, respectively. Moreover, an alternative route to the active-site gorge of the enzyme appeared to open. This structural characterization of acetylcholinesterase 'at work' is consistent with the idea that choline exits from the enzyme after catalysis either via the gorge or via an alternative 'backdoor' trajectory.
- Laboratoire de Biophysique Moléculaire, Institut de Biologie Structurale Jean-Pierre EBEL, UMR 5075 CNRS-CEA-UJF, 38027 Grenoble, France.
Organizational Affiliation: