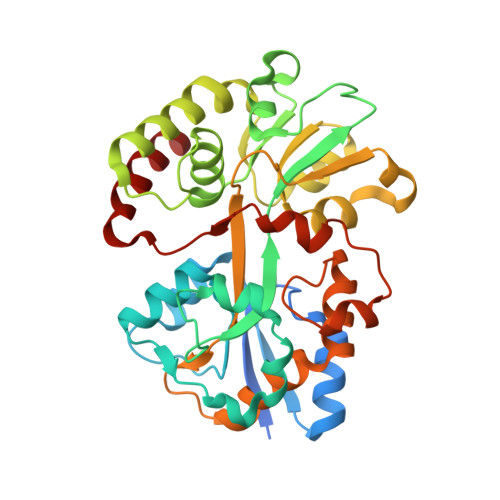
Structural and Biochemical Basis for Polyamine Binding to the Tp0655 Lipoprotein of Treponema Pallidum: Putative Role for Tp0655 (Tppotd) as a Polyamine Receptor.
Machius, M., Brautigam, C.A., Tomchick, D.R., Ward, P., Otwinowski, Z., Blevins, J.S., Deka, R.K., Norgard, M.V.(2007) J Mol Biology 373: 681
- PubMed: 17868688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2007.08.018
- Primary Citation Related Structures:
2V84 - PubMed Abstract:
Tp0655 of Treponema pallidum, the causative agent of syphilis, is predicted to be a 40 kDa membrane lipoprotein. Previous sequence analysis of Tp0655 noted its homology to polyamine-binding proteins of the bacterial PotD family, which serve as periplasmic ligand-binding proteins of ATP-binding-cassette (ABC) transport systems. Here, the 1.8 A crystal structure of Tp0655 demonstrated structural homology to Escherichia coli PotD and PotF. The latter two proteins preferentially bind spermidine and putrescine, respectively. All of these proteins contain two domains that sandwich the ligand between them. The ligand-binding site of Tp0655 can be occupied by 2-(N-morpholino)ethanesulfanoic acid, a component of the crystallization medium. To discern the polyamine binding preferences of Tp0655, the protein was subjected to isothermal titration calorimetric experiments. The titrations established that Tp0655 binds polyamines avidly, with a marked preference for putrescine (Kd=10 nM) over spermidine (Kd=430 nM), but the related compounds cadaverine and spermine did not bind. Structural comparisons and structure-based sequence analyses provide insights into how polyamine-binding proteins recognize their ligands. In particular, these comparisons allow the derivation of rules that may be used to predict the function of other members of the PotD family. The sequential, structural, and functional homology of Tp0655 to PotD and PotF prompt the conclusion that the former likely is the polyamine-binding component of an ABC-type polyamine transport system in T. pallidum. We thus rename Tp0655 as TpPotD. The ramifications of TpPotD as a polyamine-binding protein to the parasitic strategy of T. pallidum are discussed.
- Department of Biochemistry, The University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: