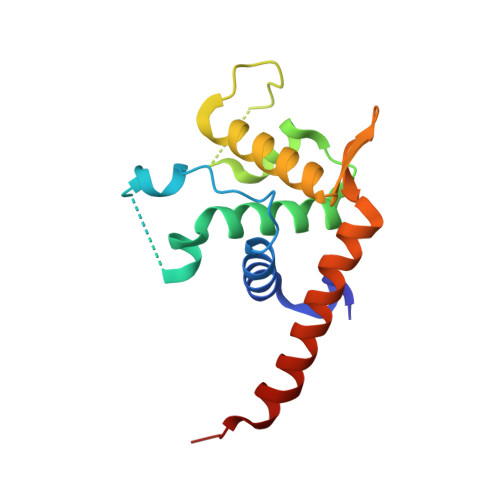
Molecular Basis of Diamond Blackfan Anemia: Structure and Function Analysis of Rps19.
Gregory, L.A., Aguissa-Toure, A.H., Pinaud, N., Legrand, P., Gleizes, P.E., Fribourg, S.(2007) Nucleic Acids Res 35: 5913
- PubMed: 17726054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkm626
- Primary Citation Related Structures:
2V7F - PubMed Abstract:
Diamond-Blackfan anemia (DBA) is a rare congenital disease linked to mutations in the ribosomal protein genes rps19, rps24 and rps17. It belongs to the emerging class of ribosomal disorders. To understand the impact of DBA mutations on RPS19 function, we have solved the crystal structure of RPS19 from Pyrococcus abyssi. The protein forms a five alpha-helix bundle organized around a central amphipathic alpha-helix, which corresponds to the DBA mutation hot spot. From the structure, we classify DBA mutations relative to their respective impact on protein folding (class I) or on surface properties (class II). Class II mutations cluster into two conserved basic patches. In vivo analysis in yeast demonstrates an essential role for class II residues in the incorporation into pre-40S ribosomal particles. This data indicate that missense mutations in DBA primarily affect the capacity of the protein to be incorporated into pre-ribosomes, thus blocking maturation of the pre-40S particles.
- INSERM U869, Institut Européen de Chimie et Biologie, 2 rue Robert Escarpit Pessac, F-33607, France.
Organizational Affiliation: