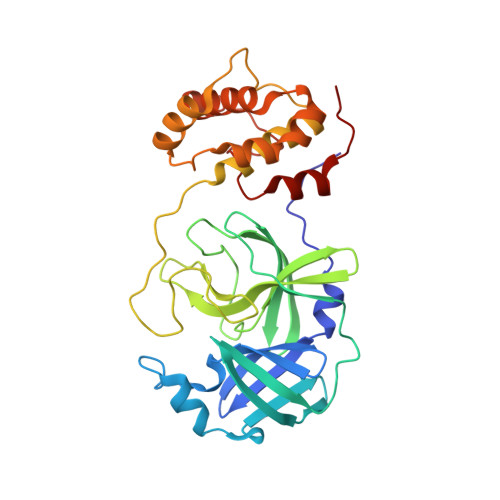
A Structural View of the Inactivation of the Sars Coronavirus Main Proteinase by Benzotriazole Esters.
Verschueren, K.H.G., Pumpor, K., Anemueller, S., Chen, S., Mesters, J.R., Hilgenfeld, R.(2008) Chem Biol 15: 597
- PubMed: 18559270 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2008.04.011
- Primary Citation Related Structures:
2V6N, 2VJ1 - PubMed Abstract:
The main proteinase (M(pro)) of the severe acute respiratory syndrome (SARS) coronavirus is a principal target for the design of anticoronaviral compounds. Benzotriazole esters have been reported as potent nonpeptidic inhibitors of the enzyme, but their exact mechanism of action remains unclear. Here we present crystal structures of SARS-CoV M(pro), the active-site cysteine of which has been acylated by benzotriazole esters that act as suicide inhibitors. In one of the structures, the thioester product has been hydrolyzed and benzoic acid is observed to bind to the hydrophobic S2 pocket. This structure also features the enzyme with a shortened N-terminal segment ("amputated N finger"). The results further the understanding of the important role of the N finger for catalysis as well as the design of benzotriazole inhibitors with improved specificity.
- Institute of Biochemistry, Center for Structural and Cell Biology in Medicine, University of Lübeck, Lübeck, Germany.
Organizational Affiliation: