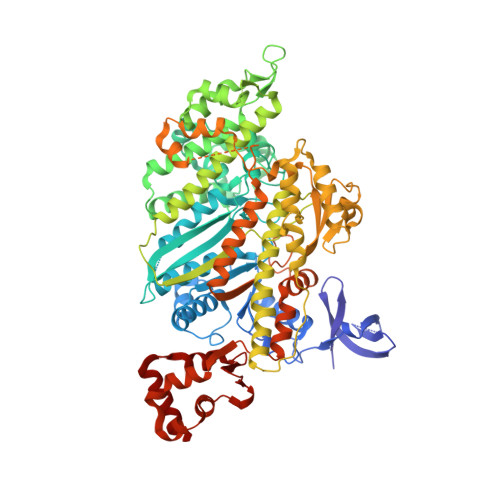
The Structural Basis for the Large Powerstroke of Myosin Vi.
Menetrey, J., Llinas, P., Mukherjea, M., Sweeney, H.L., Houdusse, A.(2007) Cell 131: 300
- PubMed: 17956731
- DOI: https://doi.org/10.1016/j.cell.2007.08.027
- Primary Citation of Related Structures:
2V26 - PubMed Abstract:
Due to a unique addition to the lever arm-positioning region (converter), class VI myosins move in the opposite direction (toward the minus-end of actin filaments) compared to other characterized myosin classes. However, the large size of the myosin VI lever arm swing (powerstroke) cannot be explained by our current view of the structural transitions that occur within the myosin motor. We have solved the crystal structure of a fragment of the myosin VI motor in the structural state that represents the starting point for movement on actin; the pre-powerstroke state. Unexpectedly, the converter itself rearranges to achieve a conformation that has not been seen for other myosins. This results in a much larger powerstroke than is achievable without the converter rearrangement. Moreover, it provides a new mechanism that could be exploited to increase the powerstroke of yet to be characterized plus-end-directed myosin classes.
- Structural Motility, Institut Curie CNRS, UMR144, 26 rue d'Ulm, 75248 Paris cedex 05, France.
Organizational Affiliation: