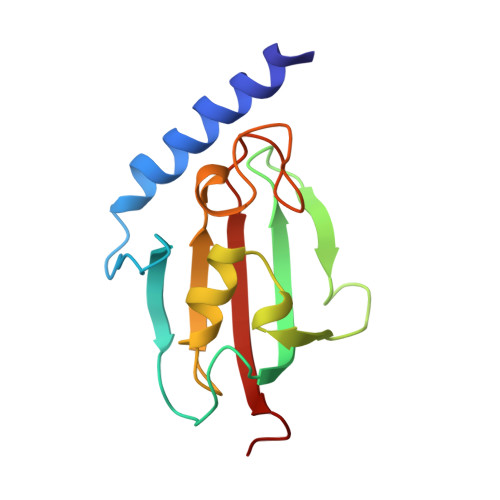
Domain Motions of the Mip Protein from Legionella Pneumophila
Horstmann, M., Ehses, P., Schweimer, K., Steinert, M., Kamphausen, T., Fischer, G., Hacker, J., Rosch, P., Faber, C.(2006) Biochemistry 45: 12303
- PubMed: 17014083 Search on PubMed
- DOI: https://doi.org/10.1021/bi060818i
- Primary Citation Related Structures:
2UZ5 - PubMed Abstract:
The homodimeric 45.6 kDa (total mass) Mip protein, a virulence factor from Legionella pneumophila, was investigated with solution NMR spectroscopy and molecular dynamics (MD) simulations. Two Mip monomers are dimerized via an N-terminal helix bundle that is connected via a long alpha-helix to a C-terminal FKBP domain in each subunit. More than 85% of the amino acids were identified in triple-resonance NMR spectra. (15)N relaxation analysis showed a bimodal distribution of R(1)/R(2) values, with the lower ratio in the N-terminal domain. Relaxation dispersion measurements confirmed that these reduced ratios did not originate from conformational exchange. Thus, two different correlation times (tau(c)) can be deduced, reflecting partly uncoupled motions of both domains. Relaxation data of a Mip(77)(-)(213) monomer mutant were similar to those observed in the dimer, corroborating that the FKBP domain, including part of the connecting helix, behaves as one dynamic entity. MD simulations (18 ns) of the Mip dimer also yielded two different correlation times for the two domains and thus confirm the independence of the domain motions. Principal component analysis of the dihedral space covariance matrix calculated from the MD trajectory suggests a flexible region in the long connecting helix that acts as a hinge between the two domains. Such motion provides a possible explanation of how Mip can bind to complex molecular components of the extracellular matrix and mediate alveolar damage and bacterial spread in the lung.
- Lehrstuhl für Experimentelle Physik 5, Universität Würzburg, Würzburg, Germany.
Organizational Affiliation: