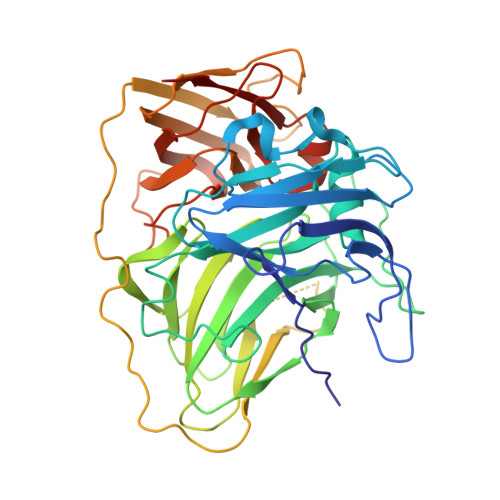
The Escherichia Coli Cell Division Protein and Model Tat Substrate Sufi (Ftsp) Localizes to the Septal Ring and Has a Multicopper Oxidase-Like Structure.
Tarry, M., Arends, S.J., Roversi, P., Piette, E., Sargent, F., Berks, B.C., Weiss, D.S., Lea, S.M.(2009) J Mol Biology 386: 504
- PubMed: 19135451 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.12.043
- Primary Citation Related Structures:
2UXT, 2UXV - PubMed Abstract:
The Escherichia coli protein SufI (FtsP) has recently been proposed to be a component of the cell division apparatus. The SufI protein is also in widespread experimental use as a model substrate in studies of the Tat (twin arginine translocation) protein transport system. We have used SufI-GFP (green fluorescent protein) fusions to show that SufI localizes to the septal ring in the dividing cell. We have also determined the structure of SufI by X-ray crystallography to a resolution of 1.9 A. SufI is structurally related to the multicopper oxidase superfamily but lacks metal cofactors. The structure of SufI suggests it serves a scaffolding rather than an enzymatic role in the septal ring and reveals regions of the protein likely to be involved in the protein-protein interactions required to assemble SufI at the septal ring.
- Department of Biochemistry, University of Oxford, UK.
Organizational Affiliation: