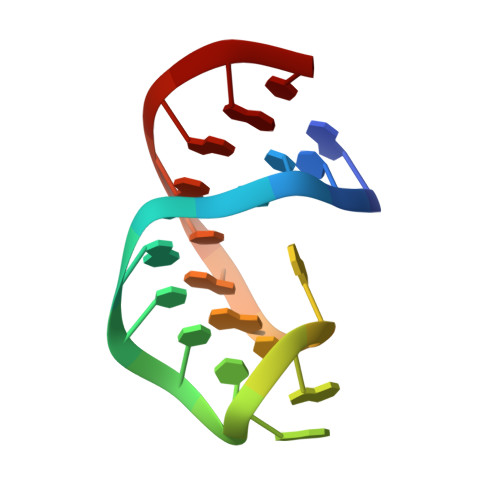
Solution structure of the tobramycin-RNA aptamer complex.
Jiang, L., Patel, D.J.(1998) Nat Struct Biol 5: 769-774
- PubMed: 9731769 Search on PubMed
- DOI: https://doi.org/10.1038/1804
- Primary Citation Related Structures:
2TOB - PubMed Abstract:
We have solved the solution structure of the aminoglycoside antibiotic tobramycin complexed with a stem-loop RNA aptamer. The 14 base loop of the RNA aptamer 'zippers up' alongside the attached stem through alignment of four mismatches and one Watson-Crick pair on complex formation. The tobramycin inserts into the deep groove centered about the mismatch pairs and is partially encapsulated between its floor and a looped out guanine base that flaps over the bound antibiotic. Several potential intermolecular hydrogen bonds between the charged NH3 groups of tobramycin and acceptor atoms on base pair edges and backbone phosphates anchor the aminoglycoside antibiotic within its sequence/structure specific RNA binding pocket.
- Cellular Biochemistry & Biophysics Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10021, USA.
Organizational Affiliation: