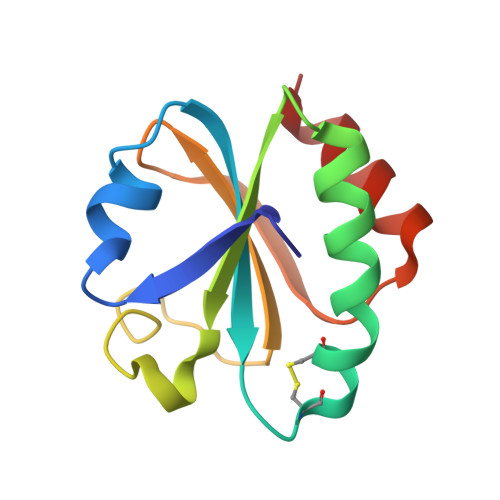
Crystal structure analysis of a mutant Escherichia coli thioredoxin in which lysine 36 is replaced by glutamic acid.
Nikkola, M., Gleason, F.K., Fuchs, J.A., Eklund, H.(1993) Biochemistry 32: 5093-5098
- PubMed: 8098620 Search on PubMed
- DOI: https://doi.org/10.1021/bi00070a017
- Primary Citation Related Structures:
2TIR - PubMed Abstract:
The structure of a mutant Escherichia coli thioredoxin with a glutamic acid substituted for a conserved lysine at position 36 adjacent to the active site has been solved using molecular replacement and refined at 2.0-A resolution to a crystallographic residual of 19.9%. The mutant was crystallized in an orthorhombic space group with one molecule in the asymmetric unit. The structure of the mutant thioredoxin shows overall good agreement with the wild-type E. coli thioredoxin. The root-mean-square deviations for all C alpha s are 0.45 and 0.79 A between the mutant structure and the two molecules in the asymmetric unit of the wild-type crystals. Structural changes are seen in several residues in the active-site region preceding the disulfide. A reverse turn of residues 29-32 changes the conformation from a type I to a type II turn. This change may be related to the loss of a hydrogen bond from Lys-36 to the main-chain carbonyl of residue 30 due to the mutation. The C alpha atom of Trp-31 has moved 1.9 A and the indole ring no longer makes hydrogen bonds to the carboxyl group of Asp-61 but instead participates in a crystal contact. The structural differences seen in the mutant thioredoxin may be influenced by the crystal packing. The substituted Glu-36 makes extensive crystal contacts. The static fluorescence of this mutant thioredoxin has a different pH dependence than the wild type.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala Biomedical Center.
Organizational Affiliation: