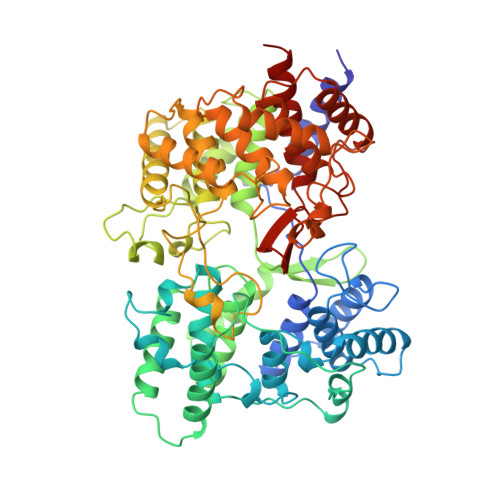
The structure of the membrane protein squalene-hopene cyclase at 2.0 A resolution.
Wendt, K.U., Lenhart, A., Schulz, G.E.(1999) J Mol Biology 286: 175-187
- PubMed: 9931258 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2470
- Primary Citation Related Structures:
2SQC, 3SQC - PubMed Abstract:
Squalene cyclases catalyze a cationic cyclization cascade, which is homologous to a key step in cholesterol biosynthesis. The structure of the enzyme from Alicyclobacillus acidocaldarius has been determined in a new crystal form at 2.0 A resolution (1 A=0.1 nm) and refined to an R-factor of 15.3 % (Rfree=18.7 %). The structure indicates how the initial protonation and the final deprotonation of squalene occur and how the transient carbocations are stabilized. The pathways of the flexible educt squalene from the membrane interior to the active center cavity and of the rigid fused-ring product hopene in the reverse direction are discussed. The enzyme contains eight so-called QW-sequence repeats that fortify the alpha/alpha-barrels by an intricate interaction network. They are unique to the known triterpene cyclases and are presumed to shield these enzymes against the released enthalpy of the highly exergonic catalyzed reaction. The enzyme is a monotopic membrane protein, the membrane-binding interactions of which are described and compared with those of two prostaglandin-H2 synthase isoenzymes, the only other structurally characterized proteins of this type. In the crystals the membrane-binding regions face each other, suggesting a micelle-type detergent structure between them.
- Institut für Organische Chemie und Biochemie, Albertstr. 21, Freiburg im Breisgau, D-79104, Germany.
Organizational Affiliation: