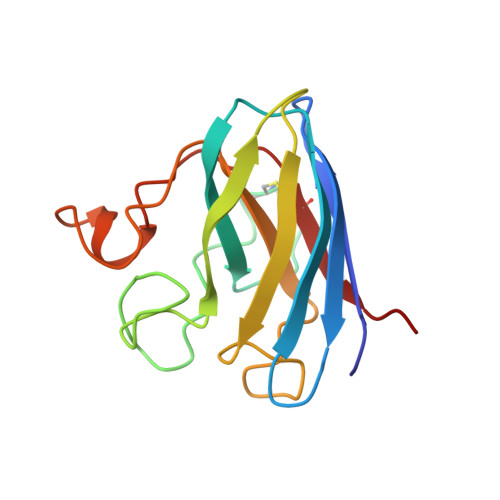
Determination and analysis of the 2 A-structure of copper, zinc superoxide dismutase.
Tainer, J.A., Getzoff, E.D., Beem, K.M., Richardson, J.S., Richardson, D.C.(1982) J Mol Biology 160: 181-217
- PubMed: 7175933 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(82)90174-7
- Primary Citation Related Structures:
2SOD