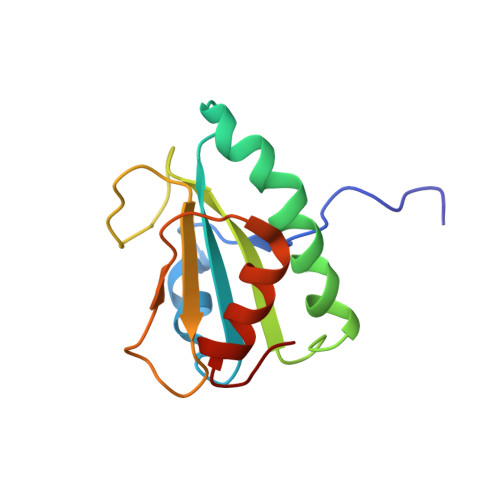
Redox-coupled structural changes of the catalytic a' domain of protein disulfide isomerase.
Inagaki, K., Satoh, T., Yagi-Utsumi, M., Le Gulluche, A.C., Anzai, T., Uekusa, Y., Kamiya, Y., Kato, K.(2015) FEBS Lett 589: 2690-2694
- PubMed: 26272828
- DOI: https://doi.org/10.1016/j.febslet.2015.07.041
- Primary Citation of Related Structures:
2RUE, 2RUF - PubMed Abstract:
Protein disulfide isomerase functions as a folding catalyst in the endoplasmic reticulum. Its b' and a' domains provide substrate-binding sites and undergo a redox-dependent domain rearrangement coupled to an open-closed structural change. Here we determined the first solution structure of the a' domain in its oxidized form and thereby demonstrate that oxidation of the a' domain induces significant conformational changes not only in the vicinity of the active site but also in the distal b'-interfacial segment. Based on these findings, we propose that this conformational transition triggers the domain segregation coupled with the exposure of the hydrophobic surface.
- Graduate School of Pharmaceutical Sciences, Nagoya City University, 3-1 Tanabe-dori, Mizuho-ku, Nagoya 467-8603, Japan; Okazaki Institute for Integrative Bioscience and Institute for Molecular Science, National Institutes of Natural Sciences, 5-1 Higashiyama, Myodaiji, Okazaki, Aichi 444-8787, Japan.
Organizational Affiliation: