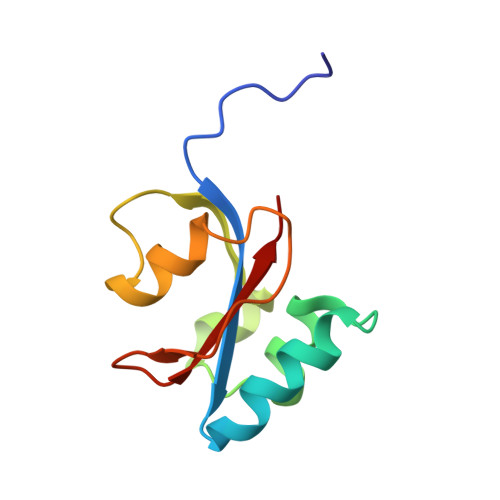
Structure and Dynamics of the First Archaeal Parvulin Reveal a New Functionally Important Loop in Parvulin-type Prolyl Isomerases
Jaremko, L., Jaremko, M., Elfaki, I., Mueller, J.W., Ejchart, A., Bayer, P., Zhukov, I.(2011) J Biological Chem 286: 6554-6565
- PubMed: 21138844
- DOI: https://doi.org/10.1074/jbc.M110.160713
- Primary Citation of Related Structures:
2RQS - PubMed Abstract:
Parvulins are a group of peptidyl-prolyl isomerases (PPIases) responsible for important biological processes in all kingdoms of life. The PinA protein from the psychrophilic archaeon Cenarchaeum symbiosum is a parvulin-like PPIase. Due to its striking similarity to the human parvulins Pin1 and Par14, PinA constitutes an interesting subject for structural and functional studies. Here, we present the first high resolution NMR structure of an archaeal parvulin, PinA, based on 1798 conformational restraints. Structure calculation yields an ensemble of 20 convergent low energy structures with a backbone r.m.s.d. value of 0.6 Å within the secondary structure elements. The overall fold of PinA comprises the β-α(3)-β-α-β(2) fold typical for all parvulin structures known so far, but with helix III being a short 3(10)-helix. A detailed comparison of this high resolution structure of the first archaeal PinA protein with bacterial and eukaryotic parvulin PPIase structures reveals an atypically large catalytic binding site. This feature provides an explanation for cold-adapted protein function. Moreover, the residues in and around 3(10)-helix III exhibit strong intramolecular dynamics on a microsecond to millisecond timescale and display structural heterogeneity within the NMR ensemble. A putative peptide ligand was found for PinA by phage display and was used for (1)H-(15)N-HSQC titrations. Again, the flexible region around 3(10)-helix III as well as residues of the peptide binding pocket showed the strongest chemical shift perturbations upon peptide binding. The local flexibility of this region also was modulated by ligand binding. A glycine and two positively charged residues are conserved in most parvulin proteins in this flexible loop region, which may be of general functional importance for parvulin-type PPIases.
- Institute of Biochemistry and Biophysics, Polish Academy of Sciences, Pawińskiego 5a, 02-106 Warsaw, Poland.
Organizational Affiliation: