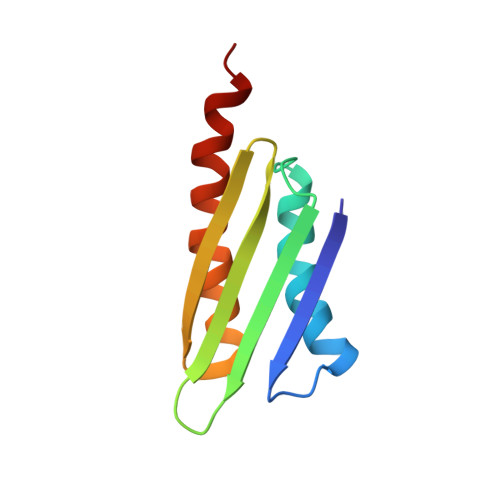
Solution structure of the E. coli ribosome hibernation promoting factor HPF: Implications for the relationship between structure and function.
Sato, A., Watanabe, T., Maki, Y., Ueta, M., Yoshida, H., Ito, Y., Wada, A., Mishima, M.(2009) Biochem Biophys Res Commun 389: 580-585
- PubMed: 19747895 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2009.09.022
- Primary Citation Related Structures:
2RQL - PubMed Abstract:
The 70S Escherichia coli ribosome dimerizes to form an inactive 100S ribosome during stationary phase, which is called "ribosome hibernation". The hibernation promoting factor HPF plays a crucial role in 100S ribosome formation. However, YfiA, a known paralog of HPF inhibits 100S formation, although it shares high sequence similarity. Here, we report the first solution structure of HPF as determined by multi-dimensional NMR. HPF adopts betaalphabetabetabetaalpha-fold and the overall structure is similar to YfiA as expected. However, detailed structure comparison based on the determined structure in this study revealed that there are remarkable differences around the C-terminal portion of helix alpha2, which is not predicted by homology modeling. Furthermore, some acidic residues conserved only in HPF are located at the rim of the common basic patch.
- Graduate School of Science and Technology, Tokyo Metropolitan University, 1-1 Minamiosawa, Hachioji, Japan.
Organizational Affiliation: