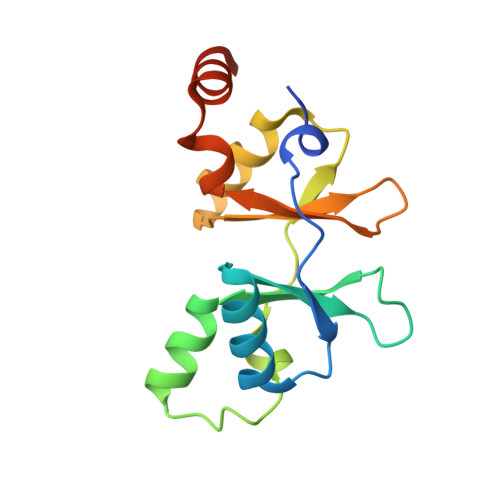
Structures and Functional Implications of an AMP-Binding Cystathionine beta-Synthase Domain Protein from a Hyperthermophilic Archaeon.
King, N.P., Lee, T.M., Sawaya, M.R., Cascio, D., Yeates, T.O.(2008) J Mol Biology 380: 181-192
- PubMed: 18513746 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2008.04.073
- Primary Citation Related Structures:
2RIF, 2RIH - PubMed Abstract:
Cystathionine beta-synthase domains are found in a myriad of proteins from organisms across the tree of life and have been hypothesized to function as regulatory modules that sense the energy charge of cells. Here we characterize the structure and stability of PAE2072, a dimeric tandem cystathionine beta-synthase domain protein from the hyperthermophilic crenarchaeon Pyrobaculum aerophilum. Crystal structures of the protein in unliganded and AMP-bound forms, determined at resolutions of 2.10 and 2.35 A, respectively, reveal remarkable conservation of key functional features seen in the gamma subunit of the eukaryotic AMP-activated protein kinase. The structures also confirm the presence of a suspected intermolecular disulfide bond between the two subunits that is shown to stabilize the protein. Our AMP-bound structure represents a first step in investigating the function of a large class of uncharacterized prokaryotic proteins. In addition, this work extends previous studies that have suggested that, in certain thermophilic microbes, disulfide bonds play a key role in stabilizing intracellular proteins and protein-protein complexes.
- Department of Chemistry and Biochemistry, University of California, Los Angeles, Los Angeles, CA 90095, USA.
Organizational Affiliation: