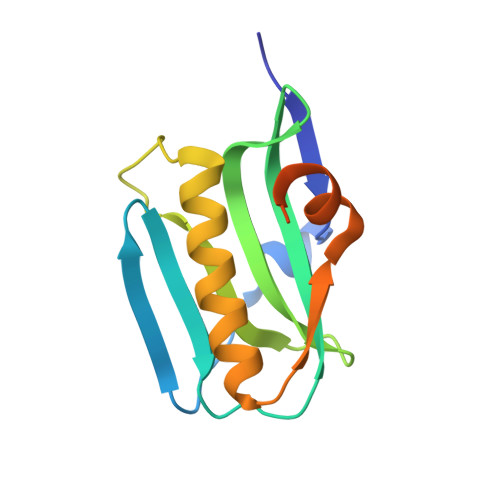
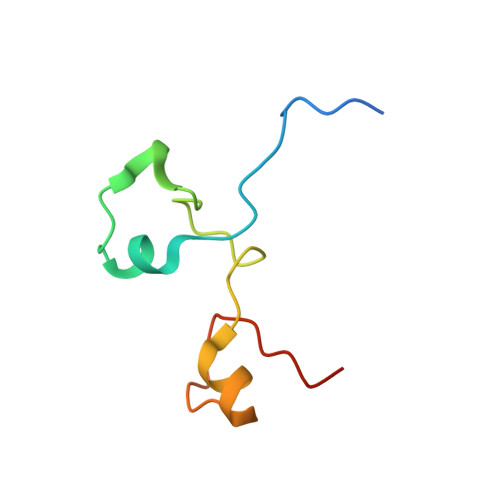
Structural basis for suppression of a host antiviral response by influenza A virus.
Das, K., Ma, L.C., Xiao, R., Radvansky, B., Aramini, J., Zhao, L., Marklund, J., Kuo, R.L., Twu, K.Y., Arnold, E., Krug, R.M., Montelione, G.T.(2008) Proc Natl Acad Sci U S A 105: 13093-13098
- PubMed: 18725644 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0805213105
- Primary Citation Related Structures:
2RHK - PubMed Abstract:
Influenza A viruses are responsible for seasonal epidemics and high mortality pandemics. A major function of the viral NS1A protein, a virulence factor, is the inhibition of the production of IFN-beta mRNA and other antiviral mRNAs. The NS1A protein of the human influenza A/Udorn/72 (Ud) virus inhibits the production of these antiviral mRNAs by binding the cellular 30-kDa subunit of the cleavage and polyadenylation specificity factor (CPSF30), which is required for the 3' end processing of all cellular pre-mRNAs. Here we report the 1.95-A resolution X-ray crystal structure of the complex formed between the second and third zinc finger domain (F2F3) of CPSF30 and the C-terminal domain of the Ud NS1A protein. The complex is a tetramer, in which each of two F2F3 molecules wraps around two NS1A effector domains that interact with each other head-to-head. This structure identifies a CPSF30 binding pocket on NS1A comprised of amino acid residues that are highly conserved among human influenza A viruses. Single amino acid changes within this binding pocket eliminate CPSF30 binding, and a recombinant Ud virus expressing an NS1A protein with such a substitution is attenuated and does not inhibit IFN-beta pre-mRNA processing. This binding pocket is a potential target for antiviral drug development. The crystal structure also reveals that two amino acids outside of this pocket, F103 and M106, which are highly conserved (>99%) among influenza A viruses isolated from humans, participate in key hydrophobic interactions with F2F3 that stabilize the complex.
- Center for Advanced Biotechnology and Medicine, Rutgers University, Piscataway, NJ 08854, USA.
Organizational Affiliation: