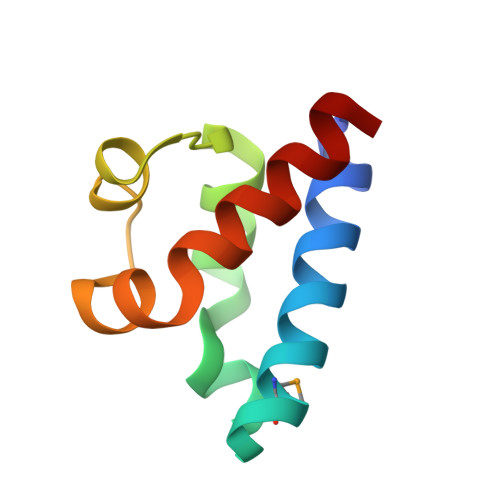
Structure and function of the regulatory C-terminal HRDC domain from Deinococcus radiodurans RecQ.
Killoran, M.P., Keck, J.L.(2008) Nucleic Acids Res 36: 3139-3149
- PubMed: 18411208 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkn143
- Primary Citation Related Structures:
2RHF - PubMed Abstract:
RecQ helicases are critical for maintaining genome integrity in organisms ranging from bacteria to humans by participating in a complex network of DNA metabolic pathways. Their diverse cellular functions require specialization and coordination of multiple protein domains that integrate catalytic functions with DNA-protein and protein-protein interactions. The RecQ helicase from Deinococcus radiodurans (DrRecQ) is unusual among RecQ family members in that it has evolved to utilize three 'Helicase and RNaseD C-terminal' (HRDC) domains to regulate its activity. In this report, we describe the high-resolution structure of the C-terminal-most HRDC domain of DrRecQ. The structure reveals unusual electrostatic surface features that distinguish it from other HRDC domains. Mutation of individual residues in these regions affects the DNA binding affinity of DrRecQ and its ability to unwind a partial duplex DNA substrate. Taken together, the results suggest the unusual electrostatic surface features of the DrRecQ HRDC domain may be important for inter-domain interactions that regulate structure-specific DNA binding and help direct DrRecQ to specific recombination/repair sites.
- Department of Biomolecular Chemistry, University of Wisconsin School of Medicine and Public Health, Madison, WI 53706-1532, USA.
Organizational Affiliation: