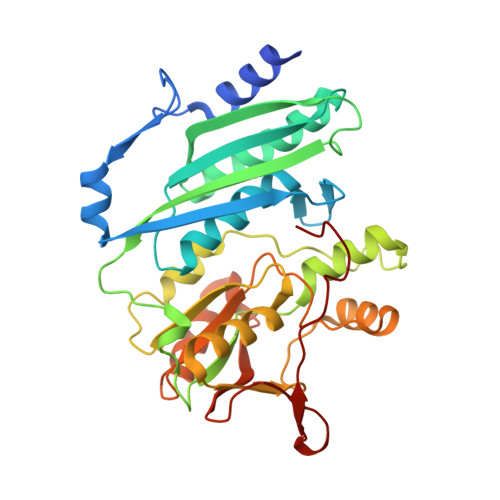
Structure of [NiFe] hydrogenase maturation protein HypE from Escherichia coli and its interaction with HypF.
Rangarajan, E.S., Asinas, A., Proteau, A., Munger, C., Baardsnes, J., Iannuzzi, P., Matte, A., Cygler, M.(2008) J Bacteriol 190: 1447-1458
- PubMed: 18065529
- DOI: https://doi.org/10.1128/JB.01610-07
- Primary Citation Related Structures:
2I6R, 2RB9 - PubMed Abstract:
Hydrogenases are enzymes involved in hydrogen metabolism, utilizing H2 as an electron source. [NiFe] hydrogenases are heterodimeric Fe-S proteins, with a large subunit containing the reaction center involving Fe and Ni metal ions and a small subunit containing one or more Fe-S clusters. Maturation of the [NiFe] hydrogenase involves assembly of nonproteinaceous ligands on the large subunit by accessory proteins encoded by the hyp operon. HypE is an essential accessory protein and participates in the synthesis of two cyano groups found in the large subunit. We report the crystal structure of Escherichia coli HypE at 2.0-A resolution. HypE exhibits a fold similar to that of PurM and ThiL and forms dimers. The C-terminal catalytically essential Cys336 is internalized at the dimer interface between the N- and C-terminal domains. A mechanism for dehydration of the thiocarbamate to the thiocyanate is proposed, involving Asp83 and Glu272. The interactions of HypE and HypF were characterized in detail by surface plasmon resonance and isothermal titration calorimetry, revealing a Kd (dissociation constant) of approximately 400 nM. The stoichiometry and molecular weights of the complex were verified by size exclusion chromatography and gel scanning densitometry. These experiments reveal that HypE and HypF associate to form a stoichiometric, hetero-oligomeric complex predominantly consisting of a [EF]2 heterotetramer which exists in a dynamic equilibrium with the EF heterodimer. The surface plasmon resonance results indicate that a conformational change occurs upon heterodimerization which facilitates formation of a productive complex as part of the carbamate transfer reaction.
- Department of Biochemistry, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: