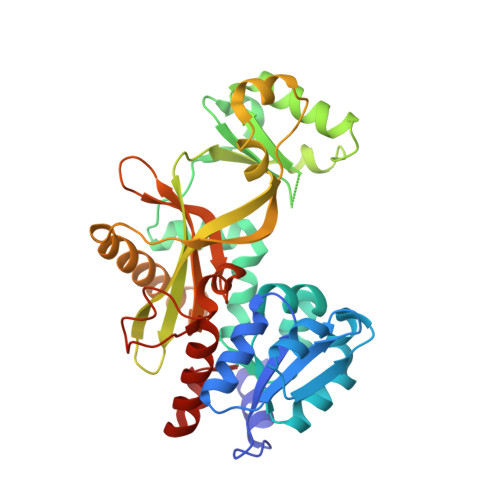
Crystal structure and function of 5-formaminoimidazole-4-carboxamide ribonucleotide synthetase from Methanocaldococcus jannaschii.
Zhang, Y., White, R.H., Ealick, S.E.(2008) Biochemistry 47: 205-217
- PubMed: 18069798
- DOI: https://doi.org/10.1021/bi701406g
- Primary Citation of Related Structures:
2R7K, 2R7L, 2R7M, 2R7N, 2R84, 2R85, 2R86, 2R87 - PubMed Abstract:
Purine biosynthesis requires 10 enzymatic steps in higher organisms, while prokaryotes require an additional enzyme for step 6. In most organisms steps 9 and 10 are catalyzed by the purH gene product, a bifunctional enzyme with both 5-formaminoimidazole-4-carboxamide ribonucleotide (FAICAR) synthase and inosine monophosphate (IMP) cyclohydrolase activity. Recently it was discovered that Archaea utilize different enzymes to catalyze steps 9 and 10. An ATP-dependent FAICAR synthetase is encoded by the purP gene, and IMP cyclohydrolase is encoded by the purO gene. We have determined the X-ray crystal structures of FAICAR synthetase from Methanocaldococcus jannaschii complexed with various ligands, including the tertiary substrate complex and product complex. The enzyme belongs to the ATP grasp superfamily and is predicted to use a formyl phosphate intermediate formed by an ATP-dependent phosphorylation. In addition, we have determined the structures of a PurP orthologue from Pyrococcus furiosus, which is functionally unclassified, in three crystal forms. With approximately 50% sequence identity, P. furiosus PurP is structurally homologous to M. jannaschii PurP. A phylogenetic analysis was performed to explore the possible role of this functionally unclassified PurP.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853-1301, USA.
Organizational Affiliation: