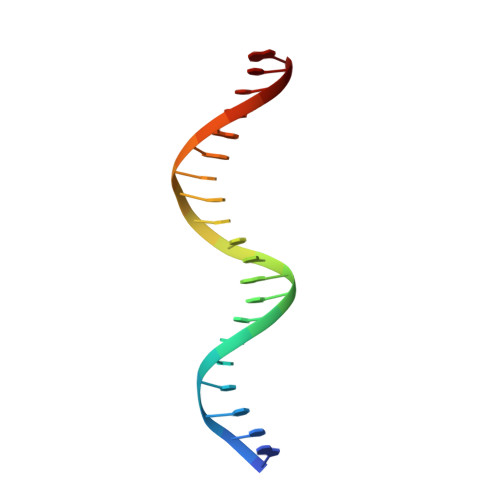
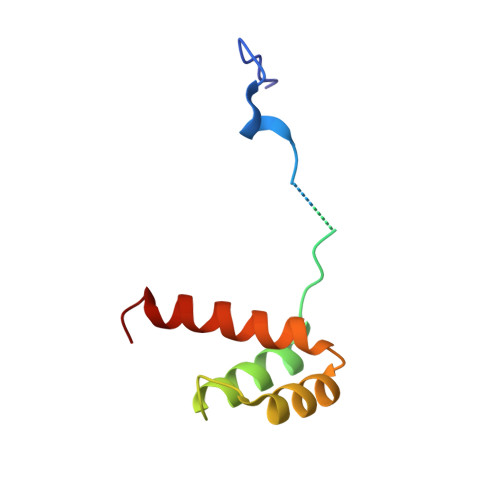
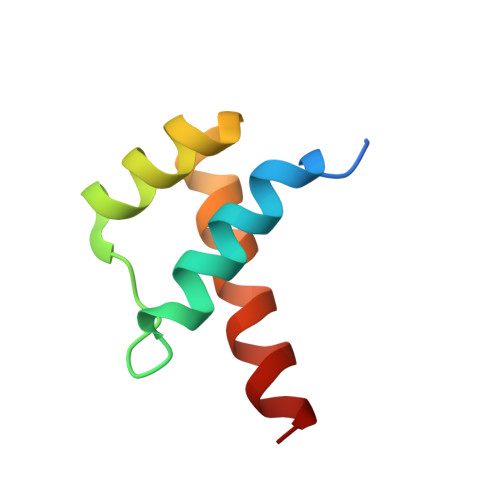
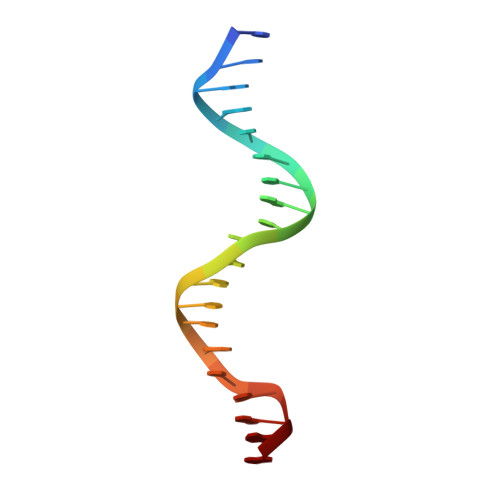
Functional specificity of a Hox protein mediated by the recognition of minor groove structure
Joshi, R., Passner, J.M., Rohs, R., Jain, R., Sosinsky, A., Crickmore, M.A., Jacob, V., Aggarwal, A.K., Honig, B., Mann, R.S.(2007) Cell 131: 530-543
- PubMed: 17981120 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2007.09.024
- Primary Citation Related Structures:
2R5Y, 2R5Z - PubMed Abstract:
The recognition of specific DNA-binding sites by transcription factors is a critical yet poorly understood step in the control of gene expression. Members of the Hox family of transcription factors bind DNA by making nearly identical major groove contacts via the recognition helices of their homeodomains. In vivo specificity, however, often depends on extended and unstructured regions that link Hox homeodomains to a DNA-bound cofactor, Extradenticle (Exd). Using a combination of structure determination, computational analysis, and in vitro and in vivo assays, we show that Hox proteins recognize specific Hox-Exd binding sites via residues located in these extended regions that insert into the minor groove but only when presented with the correct DNA sequence. Our results suggest that these residues, which are conserved in a paralog-specific manner, confer specificity by recognizing a sequence-dependent DNA structure instead of directly reading a specific DNA sequence.
- Department of Biochemistry and Molecular Biophysics, Columbia University, HHSC 1104, New York, NY 10032, USA.
Organizational Affiliation: