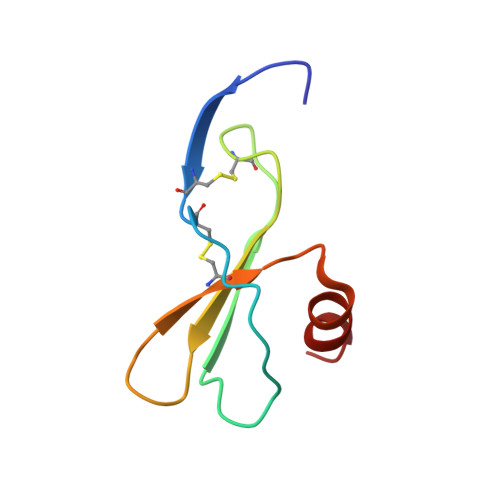
Structure of mouse IP-10, a chemokine
Jabeen, T., Leonard, P., Jamaluddin, H., Acharya, K.R.(2008) Acta Crystallogr D Biol Crystallogr 64: 611-619
- PubMed: 18560148 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444908007026
- Primary Citation Related Structures:
2R3Z - PubMed Abstract:
Interferon-gamma-inducible protein (IP-10) belongs to the CXC class of chemokines and plays a significant role in the pathophysiology of various immune and inflammatory responses. It is also a potent angiostatic factor with antifibrotic properties. The biological activities of IP-10 are exerted by interactions with the G-protein-coupled receptor CXCR3 expressed on Th1 lymphocytes. IP-10 thus forms an attractive target for structure-based rational drug design of anti-inflammatory molecules. The crystal structure of mouse IP-10 has been determined and reveals a novel tetrameric association. In the tetramer, two conventional CXC chemokine dimers are associated through their N-terminal regions to form a 12-stranded elongated beta-sheet of approximately 90 A in length. This association differs significantly from the previously studied tetramers of human IP-10, platelet factor 4 and neutrophil-activating peptide-2. In addition, heparin- and receptor-binding residues were mapped on the surface of IP-10 tetramer. Two heparin-binding sites were observed on the surface and were present at the interface of each of the two beta-sheet dimers. The structure supports the formation of higher order oligomers of IP-10, as observed in recent in vivo studies with mouse IP-10, which will have functional relevance.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, England.
Organizational Affiliation: