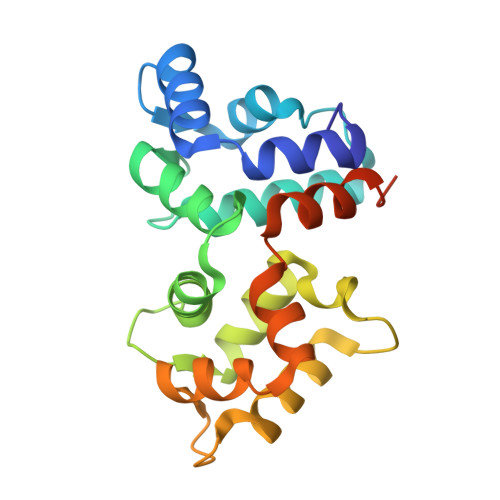
Stabilizing function for myristoyl group revealed by the crystal structure of a neuronal calcium sensor, guanylate cyclase-activating protein 1.
Stephen, R., Bereta, G., Golczak, M., Palczewski, K., Sousa, M.C.(2007) Structure 15: 1392-1402
- PubMed: 17997965 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2007.09.013
- Primary Citation Related Structures:
2R2I - PubMed Abstract:
Guanylate cyclase-activating proteins (GCAPs) are Ca(2+)-binding proteins myristoylated at the N terminus that regulate guanylate cyclases in photoreceptor cells and belong to the family of neuronal calcium sensors (NCS). Many NCS proteins display a recoverin-like "calcium-myristoyl switch" whereby the myristoyl group, buried inside the protein in the Ca(2+)-free state, becomes fully exposed upon Ca(2+) binding. Here we present a 2.0 A resolution crystal structure of myristoylated GCAP1 with Ca(2+) bound. The acyl group is buried inside Ca(2+)-bound GCAP1. This is in sharp contrast to Ca(2+)-bound recoverin, where the myristoyl group is solvent exposed. Furthermore, we provide direct evidence that the acyl group in GCAP1 remains buried in the Ca(2+)-free state and does not undergo switching. A pronounced kink in the C-terminal helix and the presence of the myristoyl group allow clustering of sequence elements crucial for GCAP1 activity.
- Department of Chemistry and Biochemistry, University of Colorado at Boulder, Boulder, CO 80309, USA.
Organizational Affiliation: