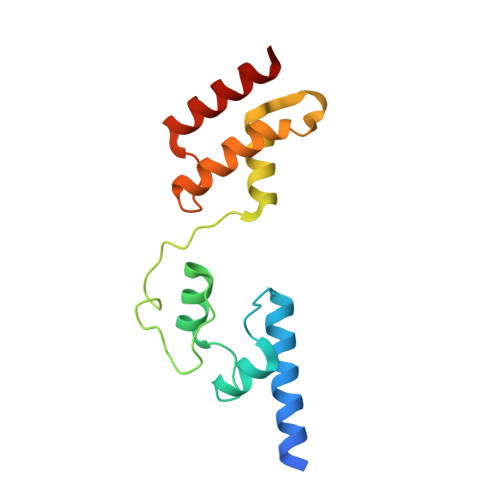
Structural insights into the multifunctional protein VP3 of birnaviruses.
Casanas, A., Navarro, A., Ferrer-Orta, C., Gonzalez, D., Rodriguez, J.F., Verdaguer, N.(2008) Structure 16: 29-37
- PubMed: 18184581 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2007.10.023
- Primary Citation Related Structures:
2R18, 2Z7J - PubMed Abstract:
Infectious bursal disease virus (IBDV), a member of the Birnaviridae family, is the causative agent of one of the most harmful poultry diseases. The IBDV genome encodes five mature proteins; of these, the multifunctional protein VP3 plays an essential role in virus morphogenesis. This protein, which interacts with the structural protein VP2, with the double-stranded RNA genome, and with the virus-encoded, RNA-dependent RNA polymerase, VP1, is involved not only in the formation of the viral capsid, but also in the recruitment of VP1 into the capsid and in the encapsidation of the viral genome. Here, we report the X-ray structure of the central region of VP3, residues 92-220, consisting of two alpha-helical domains connected by a long and flexible hinge that are organized as a dimer. Unexpectedly, the overall fold of the second VP3 domain shows significant structural similarities with different transcription regulation factors.
- Institut de Biologia Molecular de Barcelona, Consejo Superior de Investigaciones Cientificas, Parc Científic de Barcelona, Josep Samitier 1-5, Barcelona, Spain.
Organizational Affiliation: