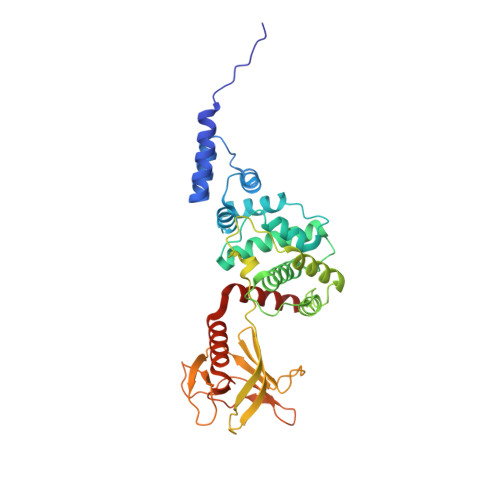
Structural Basis and Mechanism of Autoregulation in 3-Phosphoinositide-Dependent Grp1 Family Arf GTPase Exchange Factors.
DiNitto, J.P., Delprato, A., Gabe Lee, M.T., Cronin, T.C., Huang, S., Guilherme, A., Czech, M.P., Lambright, D.G.(2007) Mol Cell 28: 569-583
- PubMed: 18042453 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2007.09.017
- Primary Citation Related Structures:
2R09, 2R0D - PubMed Abstract:
Arf GTPases regulate membrane trafficking and actin dynamics. Grp1, ARNO, and Cytohesin-1 comprise a family of phosphoinositide-dependent Arf GTPase exchange factors with a Sec7-pleckstrin homology (PH) domain tandem. Here, we report that the exchange activity of the Sec7 domain is potently autoinhibited by conserved elements proximal to the PH domain. The crystal structure of the Grp1 Sec7-PH tandem reveals a pseudosubstrate mechanism of autoinhibition in which the linker region between domains and a C-terminal amphipathic helix physically block the docking sites for the switch regions of Arf GTPases. Mutations within either element result in partial or complete activation. Critical determinants of autoinhibition also contribute to insulin-stimulated plasma membrane recruitment. Autoinhibition can be largely reversed by binding of active Arf6 to Grp1 and by phosphorylation of tandem PKC sites in Cytohesin-1. These observations suggest that Grp1 family GEFs are autoregulated by mechanisms that depend on plasma membrane recruitment for activation.
- Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: