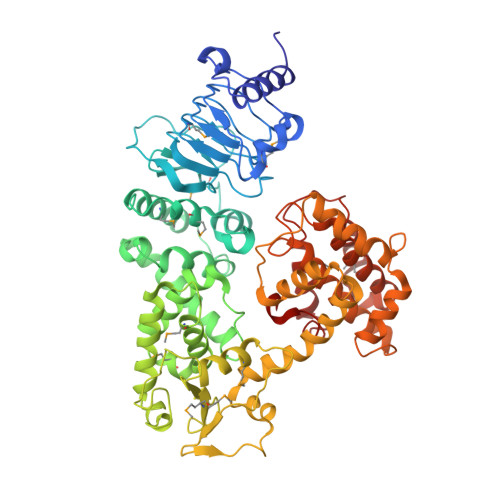
Crystal structure of SopA, a Salmonella effector protein mimicking a eukaryotic ubiquitin ligase.
Diao, J., Zhang, Y., Huibregtse, J.M., Zhou, D., Chen, J.(2008) Nat Struct Mol Biol 15: 65-70
- PubMed: 18066077
- DOI: https://doi.org/10.1038/nsmb1346
- Primary Citation Related Structures:
2QYU, 2QZA - PubMed Abstract:
Bacterial pathogens deliver virulence proteins into host cells to facilitate entry and survival. Salmonella SopA functions as an E3 ligase to manipulate the host proinflammatory response. Here we report the crystal structure of SopA in two conformations. Although it has little sequence similarity to eukaryotic HECT-domain E3s, the C-terminal half of SopA has a bilobal architecture that is reminiscent of the N- and C-lobe arrangement of HECT domains. The SopA structure also contains a putative substrate-binding domain located near the E2-binding site. The two structures of SopA differ in the relative orientations of the C lobe, indicating that SopA possesses the conformational flexibility essential for HECT E3 function. These results suggest that SopA is a unique HECT E3 ligase evolved from the coevolutionary selective pressure at the bacterium-host interface.
- Department of Biological Sciences, Purdue University, West Lafayette, Indiana 47907, USA.
Organizational Affiliation: