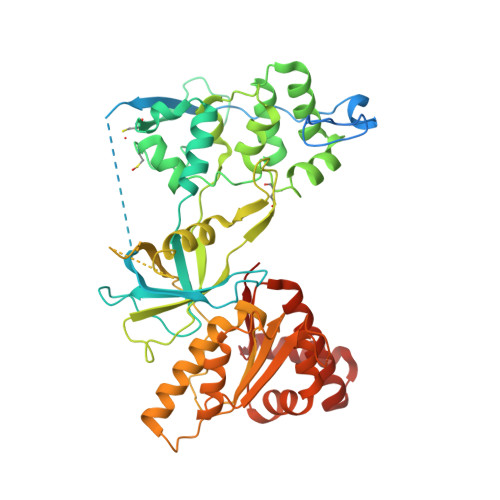
Mechanism of Coenzyme Binding to Human Methionine Synthase Reductase Revealed through the Crystal Structure of the FNR-like Module and Isothermal Titration Calorimetry
Wolthers, K.R., Lou, X., Toogood, H.S., Leys, D., Scrutton, N.S.(2007) Biochemistry 46: 11833-11844
- PubMed: 17892308 Search on PubMed
- DOI: https://doi.org/10.1021/bi701209p
- Primary Citation Related Structures:
2QTL, 2QTZ - PubMed Abstract:
Human methionine synthase reductase (MSR) is a 78 kDa flavoprotein that regenerates the active form of cobalamin-dependent methionine synthase (MS). MSR contains one FAD and one FMN cofactor per polypeptide and functions in the sequential transfer of reducing equivalents from NADPH to MS via its flavin centers. We report the 1.9 A crystal structure of the NADP+-bound FNR-like module of MSR that spans the NADP(H)-binding domain, the FAD-binding domain, the connecting domain, and part of the extended hinge region, a feature unique to MSR. The overall fold of the protein is similar to that of the corresponding domains of the related diflavin reductase enzymes cytochrome P450 reductase and neuronal nitric oxide synthase (NOS). However, the extended hinge region of MSR, which is positioned between the NADP(H)/FAD- and FMN-binding domains, is in an unexpected orientation with potential implications for the mechanism of electron transfer. Compared with related flavoproteins, there is structural variation in the NADP(H)-binding site, in particular regarding those residues that interact with the 2'-phosphate and the pyrophosphate moiety of the coenzyme. The lack of a conserved binding determinant for the 2'-phosphate does not weaken the coenzyme specificity for NADP(H) over NAD(H), which is within the range expected for the diflavin oxidoreductase family of enzymes. Isothermal titration calorimetry reveals a binding constant of 37 and 2 microM for binding of NADP+ and 2',5'-ADP, respectively, for the ligand-protein complex formed with full-length MSR or the isolated FNR module. These values are consistent with Ki values (36 microM for NADP+ and 1.4 microM for 2',5'-ADP) obtained from steady-state inhibition studies. The relatively weaker binding of NADP+ to MSR compared with other members of the diflavin oxidoreductase family might arise from unique electrostatic repulsive forces near the 5'-pyrophosphate moiety and/or increased hydrophobic stacking between Trp697 and the re face of the FAD isoalloxazine ring. Small structural permutations within the NADP(H)-binding cleft have profound affects on coenzyme binding, which likely retards catalytic turnover of the enzyme in the cell. The biological implications of an attenuated mechanism of MS reactivation by MSR on methionine and folate metabolism are discussed.
- Faculty of Life Sciences, University of Manchester, Manchester Interdisciplinary Biocentre, 131 Princess Street, Manchester M1 7DN, United Kingdom.
Organizational Affiliation: