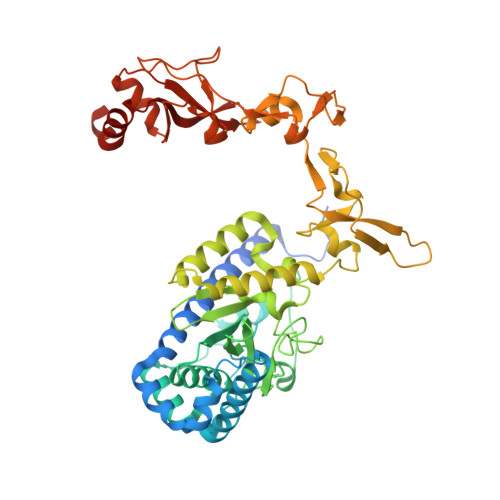
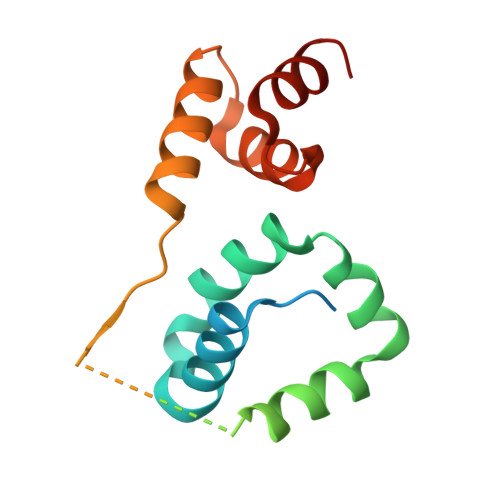
Recognition of DNA damage by the Rad4 nucleotide excision repair protein
Min, J.-H., Pavletich, N.P.(2007) Nature 449: 570-575
- PubMed: 17882165 Search on PubMed
- DOI: https://doi.org/10.1038/nature06155
- Primary Citation Related Structures:
2QSF, 2QSG, 2QSH - PubMed Abstract:
Mutations in the nucleotide excision repair (NER) pathway can cause the xeroderma pigmentosum skin cancer predisposition syndrome. NER lesions are limited to one DNA strand, but otherwise they are chemically and structurally diverse, being caused by a wide variety of genotoxic chemicals and ultraviolet radiation. The xeroderma pigmentosum C (XPC) protein has a central role in initiating global-genome NER by recognizing the lesion and recruiting downstream factors. Here we present the crystal structure of the yeast XPC orthologue Rad4 bound to DNA containing a cyclobutane pyrimidine dimer (CPD) lesion. The structure shows that Rad4 inserts a beta-hairpin through the DNA duplex, causing the two damaged base pairs to flip out of the double helix. The expelled nucleotides of the undamaged strand are recognized by Rad4, whereas the two CPD-linked nucleotides become disordered. These findings indicate that the lesions recognized by Rad4/XPC thermodynamically destabilize the Watson-Crick double helix in a manner that facilitates the flipping-out of two base pairs.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10021, USA.
Organizational Affiliation: