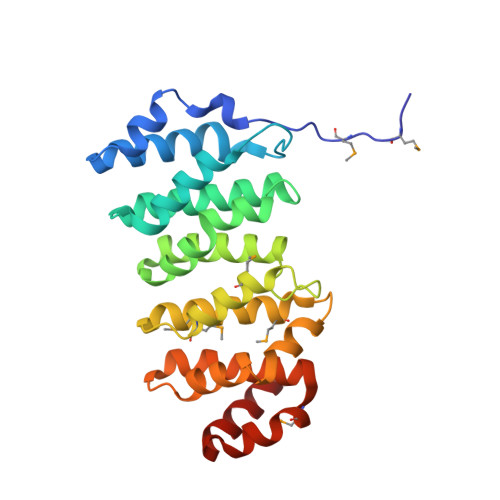
Structural Basis of Microtubule Plus End Tracking by XMAP215, CLIP-170, and EB1.
Slep, K.C., Vale, R.D.(2007) Mol Cell 27: 976-991
- PubMed: 17889670
- DOI: https://doi.org/10.1016/j.molcel.2007.07.023
- Primary Citation Related Structures:
2QJX, 2QJZ, 2QK0, 2QK1, 2QK2 - PubMed Abstract:
Microtubule plus end binding proteins (+TIPs) localize to the dynamic plus ends of microtubules, where they stimulate microtubule growth and recruit signaling molecules. Three main +TIP classes have been identified (XMAP215, EB1, and CLIP-170), but whether they act upon microtubule plus ends through a similar mechanism has not been resolved. Here, we report crystal structures of the tubulin binding domains of XMAP215 (yeast Stu2p and Drosophila Msps), EB1 (yeast Bim1p and human EB1), and CLIP-170 (human), which reveal diverse tubulin binding interfaces. Functional studies, however, reveal a common property that native or artificial dimerization of tubulin binding domains (including chemically induced heterodimers of EB1 and CLIP-170) induces tubulin nucleation/assembly in vitro and, in most cases, plus end tracking in living cells. We propose that +TIPs, although diverse in structure, share a common property of multimerizing tubulin, thus acting as polymerization chaperones that aid in subunit addition to the microtubule plus end.
- Howard Hughes Medical Institute, University of California, San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: