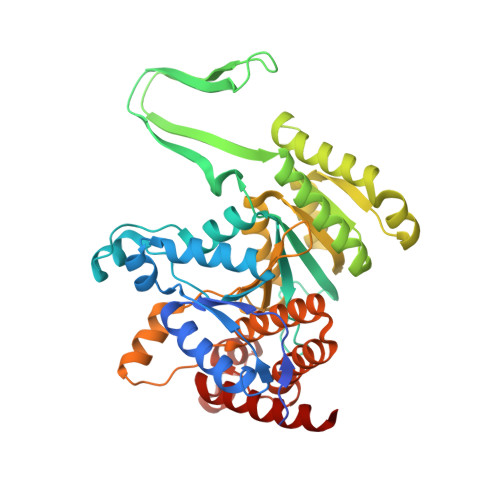
Structural studies of Saccharomyces cerevesiae mitochondrial NADP-dependent isocitrate dehydrogenase in different enzymatic states reveal substantial conformational changes during the catalytic reaction
Peng, Y.J., Zhong, C., Huang, W., Ding, J.P.(2008) Protein Sci 17: 1542-1554
- PubMed: 18552125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.035675.108
- Primary Citation Related Structures:
2QFV, 2QFW, 2QFX, 2QFY - PubMed Abstract:
Isocitrate dehydrogenases (IDHs) catalyze oxidative decarboxylation of isocitrate (ICT) into alpha-ketoglutarate (AKG). We report here the crystal structures of Saccharomyces cerevesiae mitochondrial NADP-IDH Idp1p in binary complexes with coenzyme NADP, or substrate ICT, or product AKG, and in a quaternary complex with NADPH, AKG, and Ca(2+), which represent different enzymatic states during the catalytic reaction. Analyses of these structures identify key residues involved in the binding of these ligands. Comparisons among these structures and with the previously reported structures of other NADP-IDHs reveal that eukaryotic NADP-IDHs undergo substantial conformational changes during the catalytic reaction. Binding or release of the ligands can cause significant conformational changes of the structural elements composing the active site, leading to rotation of the large domain relative to the small and clasp domains along two hinge regions (residues 118-124 and residues 284-287) while maintaining the integrity of its secondary structural elements, and thus, formation of at least three distinct overall conformations. Specifically, the enzyme adopts an open conformation when bound to NADP, a quasi-closed conformation when bound to ICT or AKG, and a fully closed conformation when bound to NADP, ICT, and Ca(2+) in the pseudo-Michaelis complex or with NADPH, AKG, and Ca(2+) in the product state. The conformational changes of eukaryotic NADP-IDHs are quite different from those of Escherichia coli NADP-IDH, for which significant conformational changes are observed only between two forms of the apo enzyme, suggesting that the catalytic mechanism of eukaryotic NADP-IDHs is more complex than that of EcIDH, and involves more fine-tuned conformational changes.
- State Key Laboratory of Molecular Biology and Research Center for Structural Biology, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China.
Organizational Affiliation: