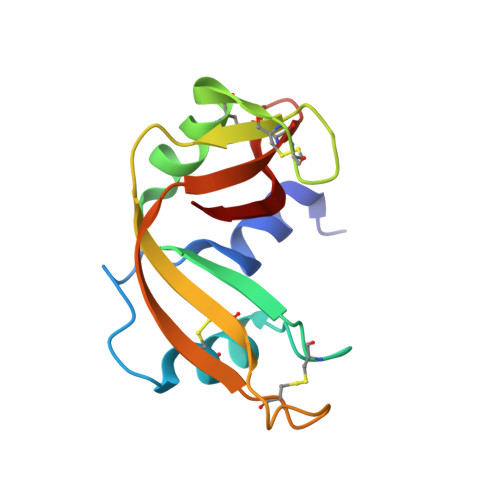
A new crystal form of bovine pancreatic RNase A in complex with 2'-deoxyguanosine-5'-monophosphate.
Larson, S.B., Day, J.S., Cudney, R., McPherson, A.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 728-733
- PubMed: 17768339 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309107039565
- Primary Citation Related Structures:
2QCA - PubMed Abstract:
The structure of bovine pancreatic RNase A has been determined in complex with 2'-deoxyguanosine-5'-monophosphate (dGMP) at 1.33 A resolution at room temperature in a previously unreported unit cell belonging to space group P3(1). There are two molecules of nucleotide per enzyme molecule, one of which lies in the active-site cleft in the productive binding mode, whilst the guanine base of the other dGMP occupies the pyrimidine-specific binding site in a nonproductive mode such that it forms hydrogen bonds to the phosphate group of the first dGMP. This is the first RNase A structure containing a guanine base in the B2 binding site. Each dGMP molecule is involved in intermolecular interactions with adjacent RNase A molecules in the lattice and the two nucleotides appear to direct the formation of the crystal lattice. Because GMP may be produced during degradation of RNA, this association could represent an inhibitor complex and thereby affect the observed enzyme kinetics.
- Department of Molecular Biology and Biochemistry, The University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: