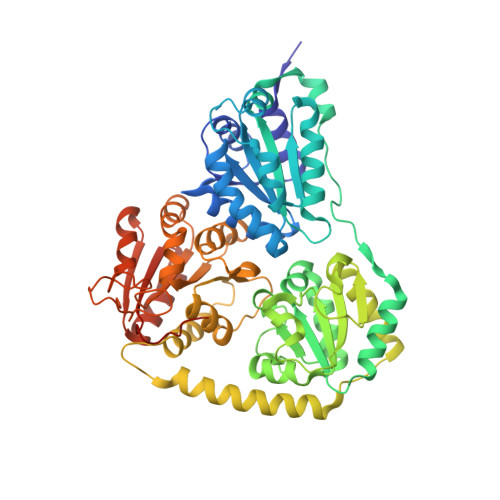
New insights into structure-function relationships of oxalyl CoA decarboxylase from Escherichia coli.
Werther, T., Zimmer, A., Wille, G., Golbik, R., Weiss, M.S., Konig, S.(2010) FEBS J 277: 2628-2640
- PubMed: 20553497 Search on PubMed
- DOI: https://doi.org/10.1111/j.1742-464X.2010.07673.x
- Primary Citation Related Structures:
2Q27, 2Q28, 2Q29 - PubMed Abstract:
The gene yfdU from Escherichia coli encodes a putative oxalyl coenzyme A decarboxylase, a thiamine diphosphate-dependent enzyme that is potentially involved in the degradation of oxalate. The enzyme has been purified to homogeneity. The kinetic constants for conversion of the substrate oxalyl coenzyme A by the enzyme in the absence and presence of the inhibitor coenzyme A, as well as in the absence and presence of the activator adenosine 5'-diphosphate, were determined using a novel continuous optical assay. The effects of these ligands on the solution and crystal structure of the enzyme were studied using small-angle X-ray scattering and X-ray crystal diffraction. Analyses of the obtained crystal structures of the enzyme in complex with the cofactor thiamine diphosphate, the activator adenosine 5'-diphosphate and the inhibitor acetyl coenzyme A, as well as the corresponding solution scattering patterns, allow comparison of the oligomer structures of the enzyme complexes under various experimental conditions, and provide insights into the architecture of substrate and effector binding sites.
- Department of Enzymology, Institute of Biochemistry & Biotechnology, Faculty for Biological Sciences, Martin Luther University Halle-Wittenberg, Halle, Germany.
Organizational Affiliation: