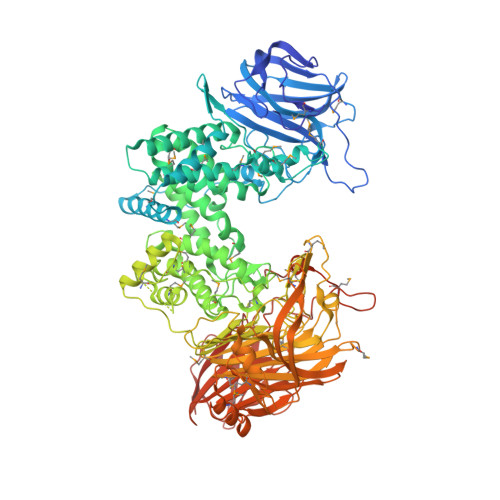
Composite active site of chondroitin lyase ABC accepting both epimers of uronic acid.
Shaya, D., Hahn, B.S., Bjerkan, T.M., Kim, W.S., Park, N.Y., Sim, J.S., Kim, Y.S., Cygler, M.(2008) Glycobiology 18: 270-277
- PubMed: 18227125
- DOI: https://doi.org/10.1093/glycob/cwn002
- Primary Citation of Related Structures:
2Q1F - PubMed Abstract:
Enzymes have evolved as catalysts with high degrees of stereospecificity. When both enantiomers are biologically important, enzymes with two different folds usually catalyze reactions with the individual enantiomers. In rare cases a single enzyme can process both enantiomers efficiently, but no molecular basis for such catalysis has been established. The family of bacterial chondroitin lyases ABC comprises such enzymes. They can degrade both chondroitin sulfate (CS) and dermatan sulfate (DS) glycosaminoglycans at the nonreducing end of either glucuronic acid (CS) or its epimer iduronic acid (DS) by a beta-elimination mechanism, which commences with the removal of the C-5 proton from the uronic acid. Two other structural folds evolved to perform these reactions in an epimer-specific fashion: (alpha/alpha)(5) for CS (chondroitin lyases AC) and beta-helix for DS (chondroitin lyases B); their catalytic mechanisms have been established at the molecular level. The structure of chondroitinase ABC from Proteus vulgaris showed surprising similarity to chondroitinase AC, including the presence of a Tyr-His-Glu-Arg catalytic tetrad, which provided a possible mechanism for CS degradation but not for DS degradation. We determined the structure of a distantly related Bacteroides thetaiotaomicron chondroitinase ABC to identify additional structurally conserved residues potentially involved in catalysis. We found a conserved cluster located approximately 12 A from the catalytic tetrad. We demonstrate that a histidine in this cluster is essential for catalysis of DS but not CS. The enzyme utilizes a single substrate-binding site while having two partially overlapping active sites catalyzing the respective reactions. The spatial separation of the two sets of residues suggests a substrate-induced conformational change that brings all catalytically essential residues close together.
- Department of Biochemistry, McGill University, Montréal, Québec, Canada.
Organizational Affiliation: