The discovery of carboline analogs as potent MAPKAP-K2 inhibitors
Wu, J.-P., Wang, J., Abeywardane, A., Andersen, D., Emmanuel, M., Gautschi, E., Goldberg, D.R., Kashem, M.A., Lukas, S., Mao, W., Martin, L., Morwick, T., Moss, N., Pargellis, C., Patel, U.R., Patnaude, L., Peet, G.W., Skow, D., Snow, R.J., Ward, Y., Werneburg, B., White, A.(2007) Bioorg Med Chem Lett 17: 4664-4669
- PubMed: 17576063 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.05.101
- Primary Citation Related Structures:
2PZY - PubMed Abstract:
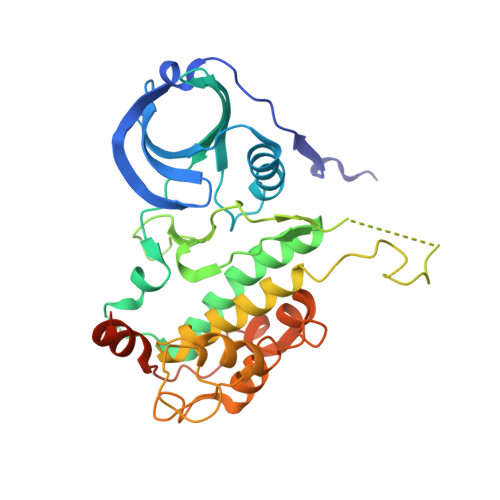
The discovery of a series of potent, carboline-based MK2 inhibitors is described. These compounds inhibit MK2 with IC50s as low as 10 nM, as measured in a DELFIA assay. An X-ray crystal structure reveals that they bind in a region near the p-loop and the hinge region of MK2a.
- Research and Development, Boehringer-Ingelheim Pharmaceuticals, 900 Ridgebury Road, Ridgefield, CT 06877, USA. jwu@rdg.boehringer-ingelheim.com
Organizational Affiliation: