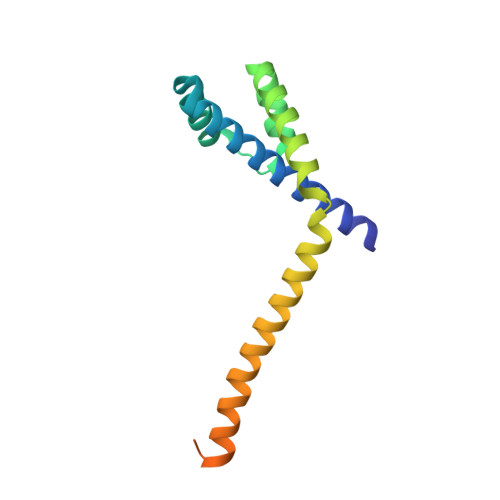
Structure of the RuBisCO chaperone RbcX from Synechocystis sp. PCC6803.
Tanaka, S., Sawaya, M.R., Kerfeld, C.A., Yeates, T.O.(2007) Acta Crystallogr D Biol Crystallogr 63: 1109-1112
- PubMed: 17881829
- DOI: https://doi.org/10.1107/S090744490704228X
- Primary Citation Related Structures:
2PY8 - PubMed Abstract:
In some cyanobacteria, the genes for the large and small subunits of the enzyme RuBisCO are separated on the bacterial chromosome by the insertion of a gene coding for a protein designated RbcX, which acts as a chaperone for RuBisCO. A recent structural study [Saschenbrecker et al. (2007), Cell, 129, 1189-1200] has shed light on the mechanism by which RbcX assists RuBisCO assembly. Here, the crystal structure of RbcX from another cyanobacterium, Synechocystis sp. PCC6803, is reported, revealing an unusually long protruding C-terminal helix, as well as a bound polyethylene glycol molecule in the protein substrate-binding site.
- UCLA Department of Chemistry and Biochemistry, 611 Charles Young Drive East, Los Angeles, CA 90095-1569, USA.
Organizational Affiliation: