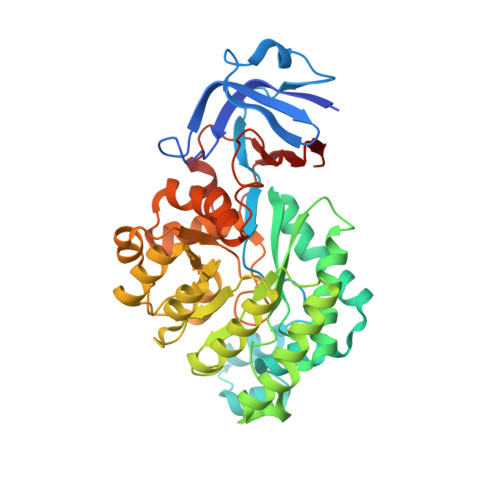
X-ray structure of imidazolonepropionase from Agrobacterium tumefaciens at 1.87 A resolution.
Tyagi, R., Kumaran, D., Burley, S.K., Swaminathan, S.(2007) Proteins 69: 652-658
- PubMed: 17640072 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21559
- Primary Citation Related Structures:
2GOK, 2PUZ - Biology Department, Brookhaven National Laboratory, Upton, New York 11973, USA.
Organizational Affiliation: