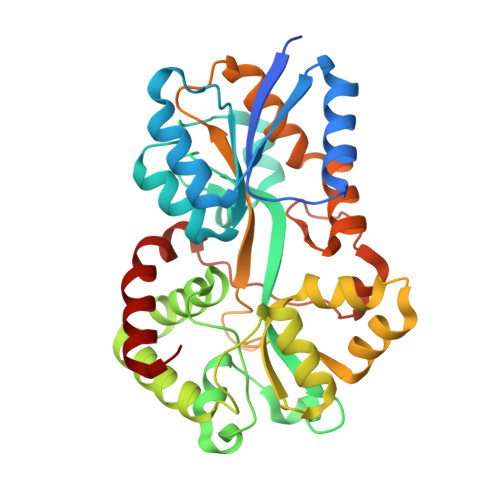
The structure of the iron-binding protein, FutA1, from Synechocystis 6803.
Koropatkin, N., Randich, A.M., Bhattacharyya-Pakrasi, M., Pakrasi, H.B., Smith, T.J.(2007) J Biological Chem 282: 27468-27477
- PubMed: 17626019 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M704136200
- Primary Citation Related Structures:
2PT1, 2PT2, 3F11 - PubMed Abstract:
Cyanobacteria account for a significant percentage of aquatic primary productivity even in areas where the concentrations of essential micronutrients are extremely low. To better understand the mechanism of iron selectivity and transport, the structure of the solute binding domain of an ATP binding cassette iron transporter, FutA1, was determined in the presence and absence of iron. The iron ion is bound within the "C-clamp" structure via four tyrosine and one histidine residues. There are extensive interactions between these ligating residues and the rest of the protein such that the conformations of the side chains remain relatively unchanged as the iron is released by the opening of the metal binding cleft. This is in stark contrast to the zinc-binding protein, ZnuA, where the domains of the metal-binding protein remain relatively fixed, whereas the ligating residues rotate out of the binding pocket upon metal release. The rotation of the domains in FutA1 is facilitated by two flexible beta-strands running along the back of the protein that act like a hinge during domain motion. This motion may require relatively little energy since total contact area between the domains is the same whether the protein is in the open or closed conformation. Consistent with the pH dependence of iron binding, the main trigger for iron release is likely the histidine in the iron-binding site. Finally, neither FutA1 nor FutA2 binds iron as a siderophore complex or in the presence of anions, and both preferentially bind ferrous over ferric ions.
- Donald Danforth Plant Science Center, Saint Louis, Missouri 63132 and.
Organizational Affiliation: