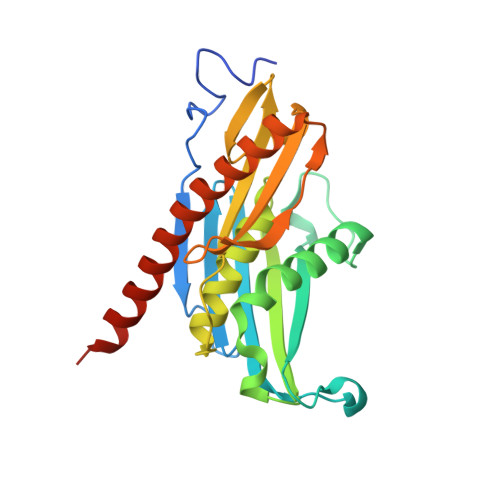
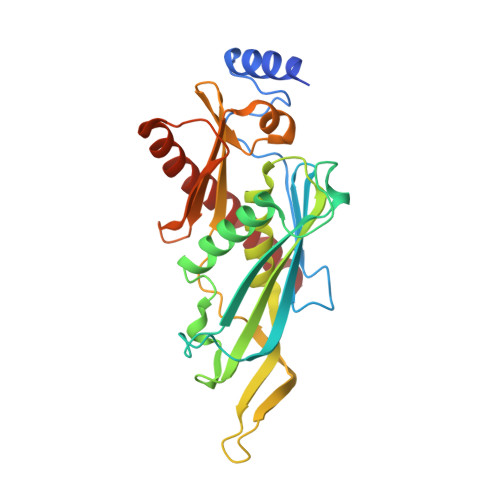
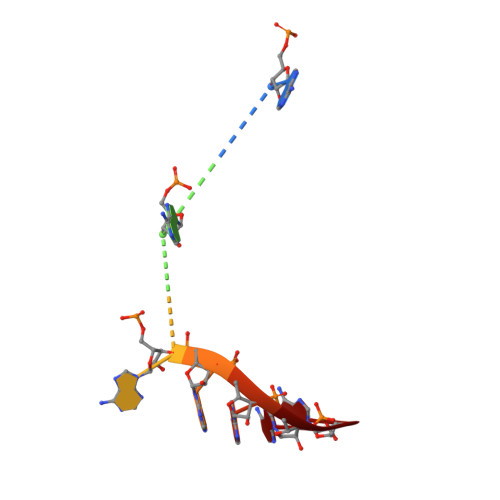
Insights into the mechanism of progressive RNA degradation by the archaeal exosome.
Navarro, M.V.A.S., Oliveira, C.C., Zanchin, N.I., Guimaraes, B.G.(2008) J Biological Chem 283: 14120-14131
- PubMed: 18353775 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M801005200
- Primary Citation Related Structures:
2PNZ, 2PO0, 2PO1, 2PO2 - PubMed Abstract:
Initially identified in yeast, the exosome has emerged as a central component of the RNA maturation and degradation machinery both in Archaea and eukaryotes. Here we describe a series of high-resolution structures of the RNase PH ring from the Pyrococcus abyssi exosome, one of them containing three 10-mer RNA strands within the exosome catalytic chamber, and report additional nucleotide interactions involving positions N5 and N7. Residues from all three Rrp41-Rrp42 heterodimers interact with a single RNA molecule, providing evidence for the functional relevance of exosome ring-like assembly in RNA processivity. Furthermore, an ADP-bound structure showed a rearrangement of nucleotide interactions at site N1, suggesting a rationale for the elimination of nucleoside diphosphate after catalysis. In combination with RNA degradation assays performed with mutants of key amino acid residues, the structural data presented here provide support for a model of exosome-mediated RNA degradation that integrates the events involving catalytic cleavage, product elimination, and RNA translocation. Finally, comparisons between the archaeal and human exosome structures provide a possible explanation for the eukaryotic exosome inability to catalyze phosphate-dependent RNA degradation.
- Brazilian Synchrotron Light Laboratory, 13083-970 Campinas, SP, Brazil. mn372@cornell.edu
Organizational Affiliation: