The Crystal Structure of PCSK9: A Regulator of Plasma LDL-Cholesterol.
Piper, D.E., Jackson, S., Liu, Q., Romanow, W.G., Shetterly, S., Thibault, S.T., Shan, B., Walker, N.P.(2007) Structure 15: 545-552
- PubMed: 17502100 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2007.04.004
- Primary Citation Related Structures:
2PMW - PubMed Abstract:
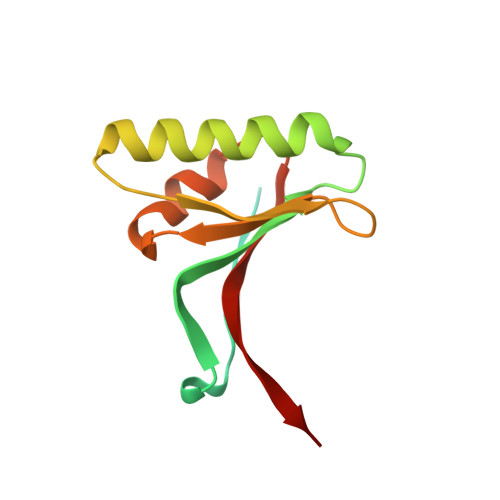
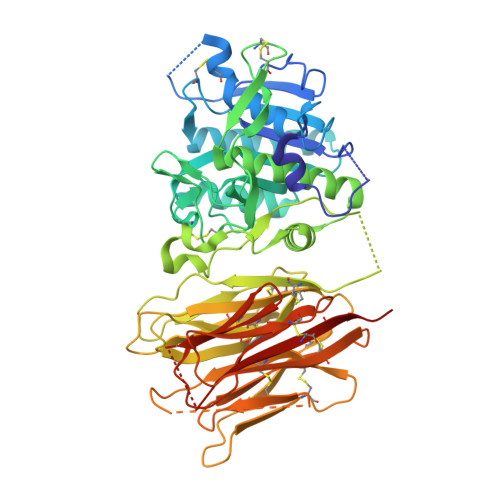
Proprotein convertase subtilisin kexin type 9 (PCSK9) has been shown to be involved in the regulation of extracellular levels of the low-density lipoprotien receptor (LDLR). Although PCSK9 is a subtilase, it has not been shown to degrade the LDLR, and its LDLR-lowering mechanism remains uncertain. Here we report the crystal structure of human PCSK9 at 2.3 A resolution. PCSK9 has subtilisin-like pro- and catalytic domains, and the stable interaction between these domains prevents access to PCSK9's catalytic site. The C-terminal domain of PCSK9 has a novel protein fold and may mediate protein-protein interactions. The structure of PCSK9 provides insight into its biochemical characteristics and biological function.
- Department of Molecular Structure, Amgen Inc., 1120 Veterans Boulevard, South San Francisco, California 94080, USA.
Organizational Affiliation: