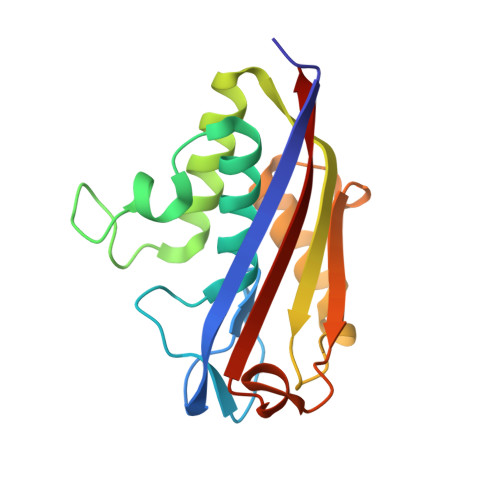
Biosynthesis of isoprenoids in plants: Structure of the 2C-methyl-D-erithrytol 2,4-cyclodiphosphate synthase from Arabidopsis thaliana. Comparison with the bacterial enzymes.
Calisto, B.M., Perez-Gil, J., Bergua, M., Querol-Audi, J., Fita, I., Imperial, S.(2007) Protein Sci 16: 2082-2088
- PubMed: 17660251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072972807
- Primary Citation Related Structures:
2PMP - PubMed Abstract:
The X-ray crystal structure of the 2C-methyl-D-erythritol 2,4-cyclodiphosphate synthase (MCS) from Arabidopsis thaliana has been solved at 2.3 A resolution in complex with a cytidine-5-monophosphate (CMP) molecule. This is the first structure determined of an MCS enzyme from a plant. Major differences between the A. thaliana and bacterial MCS structures are found in the large molecular cavity that forms between subunits and involve residues that are highly conserved among plants. In some bacterial enzymes, the corresponding cavity has been shown to be an isoprenoid diphosphate-like binding pocket, with a proposed feedback-regulatory role. Instead, in the structure from A. thaliana the cavity is unsuited for binding a diphosphate moiety, which suggests a different regulatory mechanism of MCS enzymes between bacteria and plants.
- Institut de Biologia Molecular de Barcelona-CSIC and Institut de Recerca Biomedica, Parc Cientific de Barcelona, 08028 Barcelona, Spain.
Organizational Affiliation: