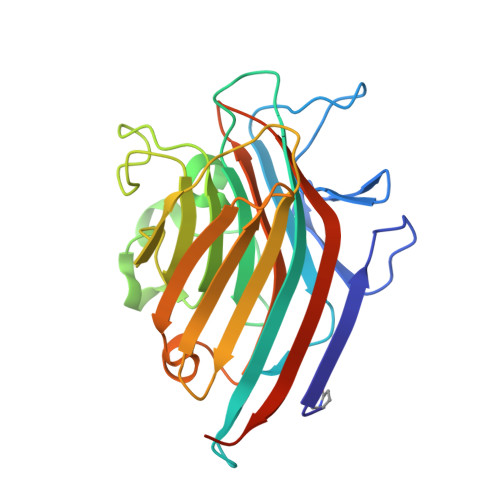
How a plant lectin recognizes high mannose oligosaccharides
Garcia-Pino, A., Buts, L., Wyns, L., Imberty, A., Loris, R.(2007) Plant Physiol 144: 1733-1741
- PubMed: 17556509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.107.100867
- Primary Citation Related Structures:
2PHF, 2PHR, 2PHT, 2PHU, 2PHW, 2PHX - PubMed Abstract:
The crystal structure of Pterocarpus angolensis seed lectin is presented in complex with a series of high mannose (Man) oligosaccharides ranging from Man-5 to Man-9. Despite that several of the nine Man residues of Man-9 have the potential to bind in the monosaccharide-binding site, all oligomannoses are bound in the same unique way, employing the tetrasaccharide sequence Manalpha(1-2)Manalpha(1-6)[Manalpha(1-3)]Manalpha(1-. Isothermal titration calorimetry titration experiments using Man-5, Man-9, and the Man-9-containing glycoprotein soybean (Glycine max) agglutinin as ligands confirm the monovalence of Man-9 and show a 4-times higher affinity for Man-9 when it is presented to P. angolensis seed lectin in a glycoprotein context.
- Laboratorium voor Ultrastructuur, Vrije Universiteit Brussel, Pleinlaan 2, B-1050 Brussel, Belgium.
Organizational Affiliation: