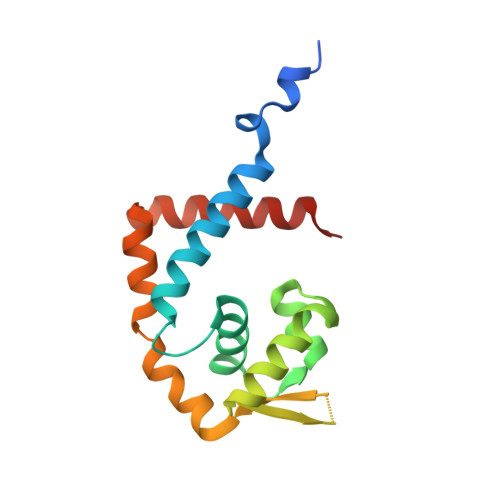
Structural Mechanism of Organic Hydroperoxide Induction of the Transcription Regulator OhrR.
Newberry, K.J., Fuangthong, M., Panmanee, W., Mongkolsuk, S., Brennan, R.G.(2007) Mol Cell 28: 652-664
- PubMed: 18042459
- DOI: https://doi.org/10.1016/j.molcel.2007.09.016
- Primary Citation of Related Structures:
2PEX, 2PFB - PubMed Abstract:
The Xanthomonas campestris transcription regulator OhrR contains a reactive cysteine residue (C22) that upon oxidation by organic hydroperoxides (OHPs) forms an intersubunit disulphide bond with residue C127'. Such modification induces the expression of a peroxidase that reduces OHPs to their less toxic alcohols. Here, we describe the structures of reduced and OHP-oxidized OhrR, visualizing the structural mechanism of OHP induction. Reduced OhrR takes a canonical MarR family fold with C22 and C127' separated by 15.5 A. OHP oxidation results in the disruption of the Y36'-C22-Y47' interaction network and dissection of helix alpha5, which then allows the 135 degrees rotation and 8.2 A translation of C127', formation of the C22-C127' disulphide bond, and alpha6-alpha6' helix-swapped reconfiguration of the dimer interface. These changes result in the 28 degrees rigid body rotations of each winged helix-turn-helix motif and DNA dissociation. Similar effector-induced rigid body rotations are expected for most MarR family members.
- Department of Biochemistry and Molecular Biology, University of Texas M.D. Anderson Cancer Center, Houston, TX 77030-4009, USA.
Organizational Affiliation: