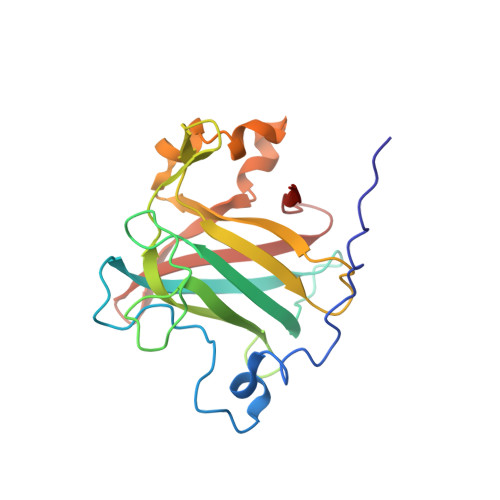
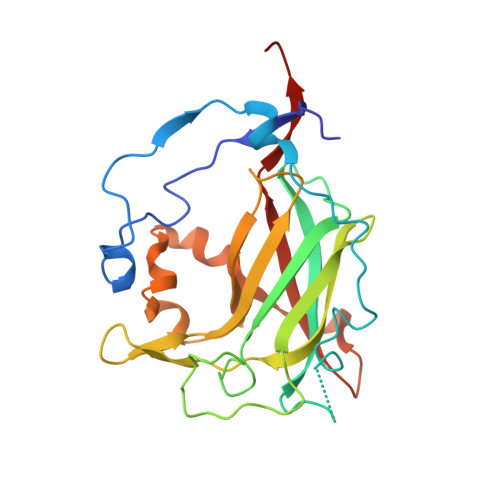
Structure of protocatechuate 3,4-dioxygenase from Pseudomonas aeruginosa at 2.15 A resolution.
Ohlendorf, D.H., Orville, A.M., Lipscomb, J.D.(1994) J Mol Biology 244: 586-608
- PubMed: 7990141 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1754
- Primary Citation Related Structures:
2PCD - PubMed Abstract:
Protocatechuate 3,4-dioxygenase catalyzes the aromatic ring cleavage of 3,4-dihydroxybenzoate by incorporating both atoms of molecular oxygen to yield beta-carboxy-cis,cis-muconate. The structure of this metalloenzyme from Pseudomonas aeruginosa (now reclassified as P. putida) has been refined to an R-factor of 0.172 to 2.15 A resolution. The structure is a highly symmetric (alpha beta Fe3+)12 aggregate with a root-mean-square (r.m.s.) difference of 0.18 A among symmetry-related atoms. The tertiary structure of the two polypeptides (alpha and beta) are highly homologous (r.m.s. difference of 1.05 A over 127 C alpha atoms), suggesting that the ancestral enzyme was originally a homodimer with two active sites. Indeed, a non-functional, vestigial active site retains many of the properties of the functional active site but does not bind iron. The coordination geometry of the non-heme iron catalytic cofactor can best be described as trigonal bipyramidal with Tyr447 (147 beta) and His462 (162 beta) serving as axial ligands, and Tyr408 (108 beta), His460 (160 beta) and Wat837 serving as equitorial ligands. The active site environment has a number of basic residues that may promote binding of the acidic substrate. Within the putative active site cavity which is located between alpha and beta chains, five approximately coplanar solvent molecules suggest a position for the planar substrate Trp449 (149 beta), Ile491 (191 beta), defined by Gly14 (14 alpha) and Pro15 (15 alpha). In this position the guanidino group of Arg457 (157 beta) would be buried by the substrate, suggesting a functional role in catalysis.
- Department of Biochemistry, University of Minnesota Medical School Minneapolis 55455-0347.
Organizational Affiliation: